 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.6G055700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 748aa MW: 82050.1 Da PI: 5.9876 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.8 | 3.4e-15 | 51 | 95 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
Seita.6G055700.1.p 51 RERWTEAEHKRFLEALKLYGRA-WQRIEEHVG-TKTAVQIRSHAQKF 95
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.99E-16 | 45 | 101 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.147 | 46 | 100 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.7E-16 | 49 | 98 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.5E-12 | 50 | 98 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.4E-8 | 51 | 91 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.5E-12 | 51 | 94 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.09E-8 | 53 | 96 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 748 aa Download sequence Send to blast |
MTSTPSDQAS SCWTIPFYFC FFFLDLEMEV NSSGEETVVK VRKPYTITKQ RERWTEAEHK 60 RFLEALKLYG RAWQRIEEHV GTKTAVQIRS HAQKFFTKLE KEAMNNGTSP GQAHDIDIPP 120 PRPKRKPNSP YPRKNGLSSE TPTKELPNDK SSKSNITLSS GNVQMVGDMS LQKFQRKEVS 180 EKGSCSEVLN LFRDAPSASF SSVNKSSSNN GVPRGVEPTK TEIRDMTTTE KNSINPTMEE 240 DVKEIDVPEI GRVNGIHVSS KCDYSNEEYL DFSMQQMKLK PKSSETTYVD KQTARTSHSL 300 AERNGATSIP VTGTDGTRPD QTGDQVGVNG SMNPCIHPML STDPKFDSSA TPQPFPHNYA 360 AFAPMMQCNC NQDTYRSFVN MSSTFSSMLV STLLSNPAIH AAARLAASYW PAAEGNTPVD 420 PTQENPAEGV QGRNLGSPPS MASIVAATVA AASAWWATQG LLPFFTPPMA FPFVPAPSAA 480 FTTADVPRPL DKDRDSPVEN AQKGCQEAQK QGQSEALRVA VSSESDGTRK GDMSLHTELK 540 ISPVQNTDAA PTTGADISDA FRNKKKQDRS SCGSNTPSSS DVEAGNVPEK EDKANDKAKQ 600 ASCSNSSAGD TNHRKFRSSG STSDSWKEVS EEGRMAFDAL FSREKLPQSF SPPQAEESKE 660 VAKEEEDEVT TVTVDLNKNA TTIDHDLDMM DEPRTSFPNE LSHLKLKSRK TGFKPYKRCS 720 VEAKENRVPA SDEVGTKRIR LESEEST* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the circadian clock. Binds to the promoter region of APRR1/TOC1 and TCP21/CHE to repress their transcription. Represses both CCA1 and itself. {ECO:0000269|PubMed:12015970, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:9657154}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
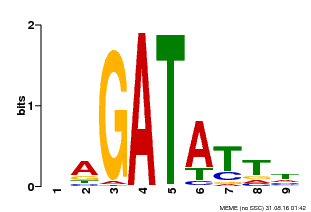
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.6G055700.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation with peak levels occurring around 1 hour after dawn. Up-regulated by APRR1/TOC1 and transiently by light treatment. Down-regulated by APRR5, APRR7 and APRR9. {ECO:0000269|PubMed:12574129, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19286557, ECO:0000269|PubMed:20233950, ECO:0000269|PubMed:9657154}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004972798.1 | 0.0 | protein LHY | ||||
| Refseq | XP_004972799.1 | 0.0 | protein LHY | ||||
| Refseq | XP_012702448.1 | 0.0 | protein LHY | ||||
| Refseq | XP_012702451.1 | 0.0 | protein LHY | ||||
| Refseq | XP_012702452.1 | 0.0 | protein LHY | ||||
| Refseq | XP_022683283.1 | 0.0 | protein LHY | ||||
| Refseq | XP_022683284.1 | 0.0 | protein LHY | ||||
| Swissprot | Q6R0H1 | 2e-42 | LHY_ARATH; Protein LHY | ||||
| TrEMBL | A0A368RIS3 | 0.0 | A0A368RIS3_SETIT; Uncharacterized protein | ||||
| STRING | Si013398m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 1e-40 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.6G055700.1.p |



