 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676775108 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 449aa MW: 48765 Da PI: 5.2727 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 124.1 | 4.6e-39 | 123 | 182 | 2 | 61 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkks 61
++k+l+cprC+s++tkfCyynny+++qPr+fCk+C+ryWt+GG++rnvPvG+grrknk+
676775108 123 PDKILPCPRCNSMETKFCYYNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNP 182
7899*****************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 7.0E-32 | 119 | 180 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 8.8E-32 | 125 | 181 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.64 | 127 | 181 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 129 | 165 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 449 aa Download sequence Send to blast |
MSDPAIKLFG KTIPLQELVD SSSTEHQDQN PVRLPDSCTV DDEEMGDSCL GEGGDDGGLH 60 DSESEKEEKD KDCGEESLRD ESSNVGTGAS SVTEKTETAK AAKTEEESSP NGTCSQEGKL 120 KKPDKILPCP RCNSMETKFC YYNNYNVNQP RHFCKKCQRY WTAGGTMRNV PVGAGRRKNK 180 NPASHYNRHV SITSAEAMQK AVSRTGDLQH PNGTNLLTFG SDSVLCESMA SGLNLAEKSM 240 LKTQTGLQEA NEGLKITVPL NPPSEEAGTI SPLPKVPCFP GPPPTWPYAW NGVSWTVVPF 300 YPPPAYWSCP GVSPGTWNSF TWMPQPTSPS GGSGPNSPTL GKHSRDESVG EPGTTLEETE 360 SLGREKSKSE RCLWVPKTLR VDDPEEAAKS SIWETLGIKK DEKADTFGAF RSSNKEKSNL 420 SEGKLSGRRP ELQANPAALS RSANFHESS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
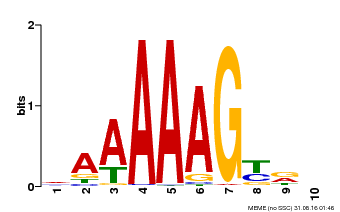
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Regulates a photoperiodic flowering response. Transcriptional repressor of 'CONSTANS' expression. The stability of CDF2 is controlled by 'GIGANTEA' and redundantly by ADO3, ADO2 and/or ADO1. {ECO:0000250, ECO:0000269|PubMed:19619493}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00046 | PBM | Transfer from AT5G39660 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676775108 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Highly expressed at the beginning of the light period, then decreases, reaching a minimum between 16 and 29 hours after dawn before rising again at the end of the day. Regulated at the protein level by ADO3 and GI pos-transcriptionally. {ECO:0000269|PubMed:19619493}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK352939 | 0.0 | AK352939.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-18-K13. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001318708.1 | 0.0 | cycling DOF factor 2 | ||||
| Refseq | NP_568567.1 | 0.0 | cycling DOF factor 2 | ||||
| Refseq | NP_851106.1 | 0.0 | cycling DOF factor 2 | ||||
| Swissprot | Q93ZL5 | 0.0 | CDF2_ARATH; Cyclic dof factor 2 | ||||
| TrEMBL | A0A178UGH8 | 0.0 | A0A178UGH8_ARATH; CDF2 | ||||
| STRING | AT5G39660.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM13795 | 18 | 25 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G39660.2 | 0.0 | cycling DOF factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



