 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676774526 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 293aa MW: 33034 Da PI: 5.0196 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 67 | 3.7e-21 | 147 | 197 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+++AeIrdp+++g r++lg+f+ta eAa+a+++a+ +l+g
676774526 147 HYRGVRQRP-WGKFAAEIRDPNKRG--SRIWLGTFDTAIEAARAYDEAAFRLRG 197
8********.**********96655..*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 5.44E-33 | 146 | 205 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 3.1E-33 | 146 | 206 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 24.368 | 147 | 205 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.8E-37 | 147 | 211 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.56E-23 | 147 | 207 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 6.6E-15 | 147 | 197 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.1E-10 | 148 | 159 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.1E-10 | 171 | 187 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 293 aa Download sequence Send to blast |
MANPSEVSAL WFIEKHLLDE VSPVATDRWT MNESAAATES SSDSSPLIFG SSSSSSAQLD 60 FSESDIKPEI IDLSTPRFID PTPFDFDLKP TYQTGNQFEP ELKTDDSQSN RKPPLKISVP 120 IKTEWIQFKP QPEVTKPSKP IVEEKKHYRG VRQRPWGKFA AEIRDPNKRG SRIWLGTFDT 180 AIEAARAYDE AAFRLRGSKA ILNFPLEVGK WKSRSEADGE KKRKRDGIDD EDVAVVEKVL 240 KTEQNVNGKE TFPFTTSNFT ELCDWDLTGF LNFPLLSPLS PHPSFGYSQL TVV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-30 | 146 | 208 | 1 | 63 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| 2gcc_A | 2e-30 | 146 | 208 | 4 | 66 | ATERF1 |
| 3gcc_A | 2e-30 | 146 | 208 | 4 | 66 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
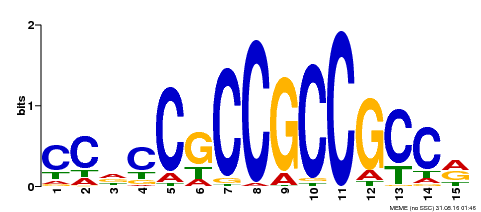
| UniProt | Acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. Involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. {ECO:0000269|PubMed:10715325, ECO:0000269|PubMed:9756931}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00549 | DAP | Transfer from AT5G47230 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676774526 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Ethylene induction is completely dependent on a functional ETHYLENE-INSENSITIVE2 (EIN2). Wounding as well as cold stress induction does not require EIN2. Transcripts accumulate strongly in cycloheximide-treated plants, a protein synthesis inhibitor. Seems to not be influenced by jasmonate, Alternaria brassicicola, exogenous abscisic acid (ABA), cold, heat, NaCl or drought stress. {ECO:0000269|PubMed:10715325}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013609112.1 | 1e-163 | PREDICTED: ethylene-responsive transcription factor 5-like | ||||
| Refseq | XP_013712994.1 | 1e-163 | ethylene-responsive transcription factor 5 | ||||
| Swissprot | O80341 | 1e-138 | EF102_ARATH; Ethylene-responsive transcription factor 5 | ||||
| TrEMBL | Q2A9W8 | 1e-162 | Q2A9W8_BRAOL; Ethylene responsive element binding factor, putative | ||||
| STRING | Bo9g069410.1 | 1e-162 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47230.1 | 1e-106 | ethylene responsive element binding factor 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



