 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676752474 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1889aa MW: 212095 Da PI: 7.1373 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 184 | 1.6e-57 | 16 | 133 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
l+e ++rwl++ ei++iL+n++k+++++e+++rp sgsl+L++rk++ryfrkDG++w+kkkdgkt++E+hekLKvg+++vl+cyYah+e n++fqrrcyw
676752474 16 LSEaQHRWLRPAEICEILQNYQKFHIASESPSRPVSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTIKEAHEKLKVGSIDVLHCYYAHGEGNENFQRRCYW 115
45559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Le++l +iv+vhylevk
676752474 116 MLEKDLMHIVFVHYLEVK 133
***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 84.786 | 12 | 138 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.0E-84 | 15 | 133 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.9E-50 | 18 | 132 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.42E-14 | 423 | 509 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 5.0E-4 | 424 | 498 | IPR013783 | Immunoglobulin-like fold |
| PROSITE profile | PS50297 | 14.477 | 606 | 678 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.5E-13 | 607 | 747 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.49E-13 | 607 | 671 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.06E-8 | 612 | 678 | No hit | No description |
| PROSITE profile | PS50088 | 8.603 | 623 | 655 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF48403 | 1.49E-13 | 701 | 746 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF52540 | 6.64E-8 | 857 | 908 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.34 | 858 | 880 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0015 | 860 | 879 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.657 | 860 | 888 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0029 | 881 | 903 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.322 | 882 | 906 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.5E-4 | 884 | 903 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF75304 | 3.4E-69 | 1042 | 1417 | IPR023631 | Amidase signature domain |
| Gene3D | G3DSA:3.90.1300.10 | 6.3E-68 | 1043 | 1419 | IPR023631 | Amidase signature domain |
| Pfam | PF01425 | 1.6E-47 | 1044 | 1416 | IPR023631 | Amidase signature domain |
| SuperFamily | SSF48452 | 3.52E-28 | 1448 | 1555 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS50293 | 21.994 | 1448 | 1549 | IPR013026 | Tetratricopeptide repeat-containing domain |
| PROSITE profile | PS50005 | 7.582 | 1448 | 1481 | IPR019734 | Tetratricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 4.7E-34 | 1448 | 1555 | IPR011990 | Tetratricopeptide-like helical domain |
| SMART | SM00028 | 0.012 | 1448 | 1481 | IPR019734 | Tetratricopeptide repeat |
| PROSITE profile | PS50005 | 7.818 | 1482 | 1515 | IPR019734 | Tetratricopeptide repeat |
| SMART | SM00028 | 7.5E-6 | 1482 | 1515 | IPR019734 | Tetratricopeptide repeat |
| Pfam | PF13181 | 0.013 | 1483 | 1515 | IPR019734 | Tetratricopeptide repeat |
| PROSITE profile | PS50005 | 9.853 | 1516 | 1549 | IPR019734 | Tetratricopeptide repeat |
| SMART | SM00028 | 0.0098 | 1516 | 1549 | IPR019734 | Tetratricopeptide repeat |
| SuperFamily | SSF53474 | 1.25E-46 | 1633 | 1885 | IPR029058 | Alpha/Beta hydrolase fold |
| Gene3D | G3DSA:3.40.50.1820 | 1.2E-47 | 1634 | 1885 | IPR029058 | Alpha/Beta hydrolase fold |
| Pfam | PF00561 | 3.4E-15 | 1639 | 1738 | IPR000073 | Alpha/beta hydrolase fold-1 |
| PRINTS | PR00111 | 5.4E-12 | 1665 | 1680 | IPR000073 | Alpha/beta hydrolase fold-1 |
| PRINTS | PR00111 | 5.4E-12 | 1709 | 1722 | IPR000073 | Alpha/beta hydrolase fold-1 |
| PRINTS | PR00111 | 5.4E-12 | 1723 | 1736 | IPR000073 | Alpha/beta hydrolase fold-1 |
| PRINTS | PR00111 | 5.4E-12 | 1830 | 1844 | IPR000073 | Alpha/beta hydrolase fold-1 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1889 aa Download sequence Send to blast |
MDVLPLYEFA DMEQLLSEAQ HRWLRPAEIC EILQNYQKFH IASESPSRPV SGSLFLFDRK 60 VLRYFRKDGH NWRKKKDGKT IKEAHEKLKV GSIDVLHCYY AHGEGNENFQ RRCYWMLEKD 120 LMHIVFVHYL EVKGSRTSIG MKENNSNSVS GTASVNIDST ASPTSTLSSL CEDADSVLAQ 180 GIVIKQVLSC EHLLNLNLEI AMFGHQLLAW ILFHRFVGTE SEKVIPENSV ENLVIRFDQP 240 CKNLLTQVQS SNTESKLVKE RTDRGGMLTA EHLIRPLQTQ LNWQIPVQDD LPLPKWPVDL 300 VPHCGMTDDT NLALFELSAQ DNFETFTSLL GSENQQSVGN SFQAPPSSME TEYIPVKKSL 360 LRHEDSLKKV DSFSRWASKE LGEMEDLQMQ SSRGDIAWTT VDCETAAAGP SLSPSLSEDQ 420 RFTIVDYWPK CAPTDAEVEV LVIGTFLLSP QEVTMYNWSC MFGEVEVPAE ILVDGVLCCH 480 APPHTAGQVP FYVTCSNRFA CSELREFDFL SGSSNKIDAA DIYGTYTKEA SLQLRFERLL 540 AHRASFHEHQ IFEDVGEKRR KISRIMLLIE EKEYLFPGIF ERDSTKQEPK ERLFREQFED 600 ELYIWLIHKV TEEGKGPNIL DEDGQGVLHF VAALGYDWAI KPILAAGVNI NFRDANGWSA 660 LHWAAFSGRF HTPMITSLTL LVGPRLHNKL TLFPVKFREE TVAVLVSLGA AAGVLTDPSP 720 EHPLGKTAAD LAYVNGHRGI SGFLAESSLT SYLEKLTMEA KENSSANSGG PKAVQTVSER 780 TAAPMSYGDV PETLSLKDSL TAVRNATQAA DRLHQVFRMQ SFQRKQLSGY DDEIGISNEL 840 AVSFAGSKTK NPGHSDVSVH SAAIHIQKKY RGWKKRKEFL LIRQRVVKIQ AHVRGHQVRK 900 QYKPIVWSVG LLEKIILRWR RKGTGLRGFK RNAIAKTVEP EPPVSSVLCP PIPQEDDYDF 960 LEKGRKQTEE RLHKALTRVK SMVQYPEARD QYRRLLTVVE GFRENEASSS SSINNRREED 1020 LVNFEDDDLI DIDSLLNDDT FMFDLKDYIT GFGSPQWKKT HEPAEKTAVV VTTLLKNGAT 1080 CVAKTIMDEM GFGITGENRH YGTPINPLMP SHVPGGCSSG SAVSVAAELV DFALGIDTTG 1140 GVRIPASFCG ILGFRPSQGT VSSIGVLPNS QSLETVGWFA RDPSVLCQVG HALLNLSAVT 1200 HKRQRSLIFA DDLFELSDIP KQKSVHVVRK AIENLSGYPA PKHMNVGQYV ASNVPSLAEF 1260 CEKSGKSQDS ASILKALSSV MLSVQRHEFK TNHEEWSQTC RSFLGPRFSN DVVTALKSRT 1320 ENIKSLYRVK TEMRATIQNL LKEDGILVIP TVADPPLKLN TKNKNAMNEF LDRNYALASI 1380 ASMSGCCQVT IPLGKHGDNP ICVSFLAYYG GDKFLLDTIL DVYASLQDQA DIASSLAPVA 1440 DSNGMEASDL MKEKGNAAYK GRQWNKAVSC YTEAIKLNGA NATYFCNRAA AYLELGRFQQ 1500 AEEDCTEAVL IDKKNVKAYL RRGTARESLI RYKEAAADFR HALVLEPQNK TAKNAEKRLR 1560 KLMITWSDVW VFSAHLLPNR VLYRITISHS FTASRDWLFR QSFANAGLRS VTTDLSHGNS 1620 IASTAMHCWI PKYPNRSKPN LLLVHGFGAN AMWQYGEHLR AFTGRFNVYV PDLLFFGLSS 1680 TSEPNRSESF QARCLMRLME AHGVHRMNIV GISYGGFVGY SLAAQFPEKV EKLVLCCAGV 1740 CLEEKDMEDG LFKVPNLDEA TDILIPQTPE KLKELIRFSF VKPIKGVPSF FLWDFIDVMC 1800 TEFVEEKRDL IKSILKDRRL SDLPRIKQKS LIIWGEEDQI FPLELGYRLK RHIGENAEMV 1860 VIKKAGHAVN LEKSKEFLKH LKSFLIDSL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6hpg_A | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| 6hpg_B | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| 6hpg_C | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| 6hpg_D | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| 6hpg_E | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| 6hpg_F | 2e-64 | 1445 | 1561 | 3 | 119 | Outer envelope protein 64, mitochondrial |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
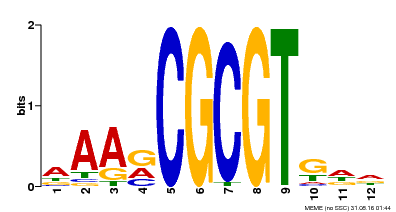
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676752474 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC189313 | 0.0 | AC189313.2 Brassica rapa subsp. pekinensis cultivar Inbred line 'Chiifu' clone KBrB034N10, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024011817.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X3 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | V4N5T0 | 0.0 | V4N5T0_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006399418.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.3 | 0.0 | ethylene induced calmodulin binding protein | ||||



