 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676730938 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 339aa MW: 37392.1 Da PI: 7.0038 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 103.5 | 3.6e-32 | 73 | 192 | 2 | 103 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec..eaesssssasnsssg........... 88
g+kdrhsk++T +g+RdRRvRls+++a++++dLq++LG+d++sk+++WLl +ak++i+el+ ++ s++++ ++++s + + + +
676730938 73 FGGKDRHSKVCTLRGLRDRRVRLSVPTAIQLYDLQERLGVDQPSKAVDWLLDAAKEEIDELPPLPVSPETFGlfNHHQSFLNLGPRTGQdptqlgfking 172
689*************************************************************777774445544444444442222234444455555 PP
TCP 89 ....kaaksaa.kskksqks 103
+a+++++ +++++++
676730938 173 cveeSASTTTTsREENNNER 192
55432222222122222222 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 31.314 | 75 | 133 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 1.1E-28 | 75 | 151 | IPR005333 | Transcription factor, TCP |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0031347 | Biological Process | regulation of defense response | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0009507 | Cellular Component | chloroplast | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MSTVPWKDTN DDVSGGTKTR REGEAEEDAE EAVAVSTSGK TVITKPPTSI SSSSSSSWMK 60 SKDPRIVRVS RAFGGKDRHS KVCTLRGLRD RRVRLSVPTA IQLYDLQERL GVDQPSKAVD 120 WLLDAAKEEI DELPPLPVSP ETFGLFNHHQ SFLNLGPRTG QDPTQLGFKI NGCVEESAST 180 TTTSREENNN ERGEKDVAFA NNHHIGSYGI YHNMEHHYHQ QHSSFQADYH HHQHQLHSLV 240 PFPSQFLVYP MTASSTTTTI QSLFPSSSSA GSGNMGTTDP RQMVGHFQMP LMGSSSSSAS 300 QNISNLYSLL HGSSSNNDSS NRMSSVQFNR TSRGRDNPM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 3e-19 | 80 | 134 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 3e-19 | 80 | 134 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164). Binds to the 3'-ACC-5' repeats in the light-responsive promoter (LRP) of psbD, and activates its transcription. Participates in ovule develpment (PubMed:25378179). {ECO:0000269|PubMed:11161017, ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:25378179}. | |||||
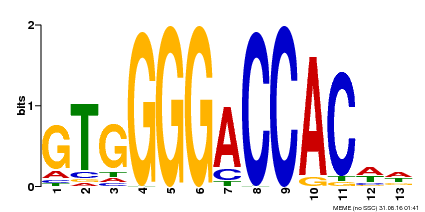
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00326 | DAP | Transfer from AT3G02150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676730938 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009134647.1 | 1e-176 | PREDICTED: transcription factor TCP13-like isoform X1 | ||||
| Refseq | XP_009134648.1 | 1e-176 | PREDICTED: transcription factor TCP13-like isoform X1 | ||||
| Refseq | XP_009134649.1 | 1e-176 | PREDICTED: transcription factor TCP13-like isoform X1 | ||||
| Refseq | XP_018513324.1 | 1e-176 | PREDICTED: transcription factor TCP13-like isoform X1 | ||||
| Swissprot | Q9S7W5 | 1e-175 | TCP13_ARATH; Transcription factor TCP13 | ||||
| TrEMBL | A0A178VKM0 | 1e-174 | A0A178VKM0_ARATH; TFPD | ||||
| TrEMBL | A0A398A2Q5 | 1e-174 | A0A398A2Q5_BRACM; Uncharacterized protein | ||||
| STRING | AT3G02150.2 | 1e-174 | (Arabidopsis thaliana) | ||||
| STRING | Bo3g055460.1 | 1e-174 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9134 | 26 | 37 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G02150.2 | 1e-137 | plastid transcription factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



