 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676728310 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 628aa MW: 68491.6 Da PI: 4.7802 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.3 | 1.1e-12 | 452 | 497 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
676728310 452 NHVEAERQRREKLNQRFYSLRAVVPNV-----SKMDKASLLGDAISYINEL 497
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.2E-55 | 69 | 270 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.282 | 448 | 497 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.57E-18 | 451 | 520 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.28E-14 | 451 | 502 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 7.9E-19 | 452 | 517 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 4.1E-10 | 452 | 497 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.3E-17 | 454 | 503 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043425 | Molecular Function | bHLH transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 628 aa Download sequence Send to blast |
MSPTNVQLTD YHLNQTTNGT NLWSTEDDTS EMEAFIGSSD HSSLWPQPPV TPPPHSLPPA 60 QSQATEDTLQ QRLQALIEGA RESWTYAVFW QLSYDFAESG GNSNRVLLGW GDGYYKGEEV 120 RKTRKKKSKP VSAAEQEHRR RVIRELNSLI SGGGGTVNNG GSGSGGGGGS SDEAGDEEVT 180 DTEWFFLVSM TQSFTDGSGL PGQAYSNSQT IWLSGSNALA GSSCQRARQG QVYGLETMVC 240 IPVENGVVEL GSLEVIHQSS DLVDKVNSFF SFNNGGGGGG GESGSWTFNL NPDQGENDPA 300 MWINEPGMEP GLVAPAGNNN STSESDSQQI SKLCNGSSVE NPSKQQNPQI SSSGFENQPK 360 VSGEVVSFKN GGNRFVEEDS NKKRCLVSDN EEAMLSFTSV LPRTTKSGDS NHSDLEPSVA 420 KEAESDRIVV EPEKKPRKRG RKPANGREEP LNHVEAERQR REKLNQRFYS LRAVVPNVSK 480 MDKASLLGDA ISYINELKAK LQKAESDKQE LQKQIDGMSK EVANGNYVKS SVKEQKCSDQ 540 DSGLLIELEI DVKIIGWDAM IRVQCSKKNH PGAKFMEALK ELELEVNHAS LSVVNDFMIQ 600 QATVKMGNQF FTQDQLKATL IEKVGEGT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rru_A | 6e-85 | 17 | 266 | 1 | 227 | Transcription factor MYC3 |
| 4ywc_A | 6e-85 | 17 | 266 | 1 | 227 | Transcription factor MYC3 |
| 4ywc_B | 6e-85 | 17 | 266 | 1 | 227 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 433 | 441 | KKPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in jasmonic acid (JA) gene regulation. With MYC2 and MYC3, controls additively subsets of JA-dependent responses. Can form complexes with all known glucosinolate-related MYBs to regulate glucosinolate biosynthesis. Binds to the G-box (5'-CACGTG-3') of promoters. Activates multiple TIFY/JAZ promoters. {ECO:0000269|PubMed:21321051, ECO:0000269|PubMed:21335373, ECO:0000269|PubMed:23943862}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00086 | PBM | Transfer from AT4G17880 | Download |
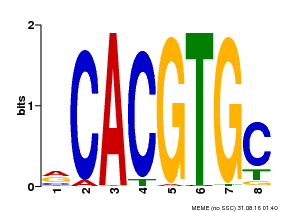
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676728310 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. Not induced by jasmonic acid. {ECO:0000269|PubMed:12679534, ECO:0000269|PubMed:21335373}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC183492 | 0.0 | AC183492.1 Brassica oleracea Contig G, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006283348.1 | 0.0 | transcription factor MYC4 | ||||
| Swissprot | O49687 | 0.0 | MYC4_ARATH; Transcription factor MYC4 | ||||
| TrEMBL | R0F4A8 | 0.0 | R0F4A8_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006283348.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1265 | 28 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G17880.1 | 0.0 | bHLH family protein | ||||



