 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011100396.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 890aa MW: 102122 Da PI: 7.405 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 86.1 | 5.5e-27 | 133 | 235 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdgkwevtklel 83
fY+eYA++vGF++ +++s++sk+++e+++++f Cs++g+++e +k+ +e++ +ra +t+Cka+++vk+++dgkw ++++e+
XP_011100396.1 133 FYQEYARSVGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKSlnrprsrqgsnqdAENATGRRACAKTDCKASMHVKRRSDGKWIIHRFEK 227
9*****************************************999999***999988776677779***************************** PP
FAR1 84 eHnHelap 91
eHnHel p
XP_011100396.1 228 EHNHELLP 235
*****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 6.1E-25 | 133 | 235 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.7E-27 | 333 | 425 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.458 | 613 | 649 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 3.2E-4 | 615 | 646 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 8.3E-8 | 624 | 651 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 890 aa Download sequence Send to blast |
MWWQLFFTFL FFTSNYQIIY VAFWRKGCCR TVFCRREKRT VVTMDIDLRL PSGGHDKEVE 60 EEPNGIVNML DGEEKPLNVD GVGGSMGDVE EKLQIEDAED VNSPIHDIDF KDVTILEPLP 120 GMEFGSHGDA YAFYQEYARS VGFNTAIQNS RRSKTSREFI DAKFACSRYG TKREYEKSLN 180 RPRSRQGSNQ DAENATGRRA CAKTDCKASM HVKRRSDGKW IIHRFEKEHN HELLPAQAVS 240 EQTRRMYAAM ARQFAEYKSA VGLKHDSRSH FEKGRNMAMD AGEANMLLDF FVQMQSLNSN 300 FFYAVDVGED QRLKNLLWID AKSRHDYPNF SDVVSFDTSY VRNKYKMPLA LFVGVNQHYQ 360 FMLLGCALVS DESEATFSWV MQTWLKAMGG QAPKIIITDQ DKVMKSVTAD AFPSTLHFFC 420 LWNIMGKVSE TLNHVIKQNE NFMSKFEKCV YRSWTDDEFD KRWHKLVNRF GLKENELMQS 480 LYEDRKKWVP NFMKDGFFAG MSTGQRSESV NSFFDKYVHK KTTMQEFIKQ YEAILQDRYE 540 EEAKASSDTW NKQPSLKSPS PFEKHVAGLY THAVFRKFQV EVLGAVACIP KREEQVDATI 600 TFKVQDFERN QEFIVTLNEL KSEVSCICRL FEFKGFLCRH AMIVLQICGI SNIPSQYILK 660 RWTKDAKSRY PMGEGSEQVQ SRLQRYNDLC QRAIKLGEEG SFSQESYNLT LRALDDAFET 720 CLNANNSCKN LLEAGPSSSP ALLCIEEDIQ SGNLSKTNKK KSSTKKRKVN MEPDVITVGT 780 PESLQQMEKL NSRPVNLDGF FGAQQGVQGM VQLNLMAPTR DNYYGNQQTI QGLGQLNSIA 840 PTHDGYYGTQ PAIHGLGQMD FFRTPSFGYG IREDPNVRSA QLHDDAPRHA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
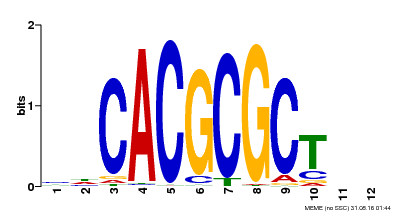
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011100396.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011100396.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2G9GLI3 | 0.0 | A0A2G9GLI3_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.M00094.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



