 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011095220.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 375aa MW: 42384 Da PI: 8.2502 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102 | 3.8e-32 | 61 | 114 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+++AtPk +l+lm++kgL ++hvkSHLQ+YR+
XP_011095220.1 61 PRLRWTPDLHMRFVHAVERLGGQDRATPKLVLQLMNIKGLNIAHVKSHLQMYRS 114
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-30 | 57 | 115 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.122 | 57 | 117 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.06E-14 | 59 | 115 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.9E-22 | 61 | 116 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.4E-8 | 62 | 113 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MDSGEKECSK TSPSEKNDDE DESQGKDESC TVKDGGSSSN STVEESEKKP SVRPYVRSKM 60 PRLRWTPDLH MRFVHAVERL GGQDRATPKL VLQLMNIKGL NIAHVKSHLQ MYRSKKIDES 120 NQGLMDHRLF MEGGDRNVYN LSQLPLLPSF NQRQNYSNLR YGDPSWSPHG KWMHNSAPGQ 180 NTISHKTRPG FYDTITERLI GSHFYRPSAN LDLWSRTSSH TDGSTWRRNE DFKEVYRQEP 240 WLGQSRQNPI QLNSSKLIQN GVQEQRISSD AAKSNPPISS AGQTMRDTAR LKRKASGSDD 300 KLDLNLSLAL ESRNESEQKG TLITDDDDEH LSLSLFTSRP SSSNLIKKLK EDDTVVHGTD 360 QQNAIRGAST LDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 5e-17 | 62 | 116 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-17 | 62 | 116 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-17 | 62 | 116 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-17 | 62 | 116 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
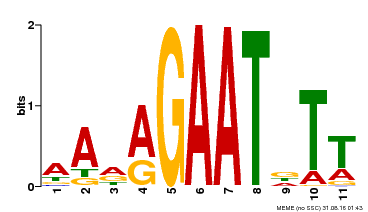
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011095220.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011095220.1 | 0.0 | uncharacterized protein LOC105174731 | ||||
| TrEMBL | A0A2G9H772 | 1e-143 | A0A2G9H772_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.J00123.1.p | 1e-118 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4382 | 21 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 3e-58 | G2-like family protein | ||||



