 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011092424.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1021aa MW: 114178 Da PI: 5.7574 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 183.7 | 2.1e-57 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqr 96
l+ ++rwl++ ei++iL n+ek++++ e++++p sgs++L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v++l+cyYah+e+n++fqr
XP_011092424.1 20 LEAQHRWLRPAEICEILRNYEKFHISPEAPNKPVSGSVFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDMLHCYYAHGEDNENFQR 114
5679******************************************************************************************* PP
CG-1 97 rcywlLeeelekivlvhylevk 118
r+ywlLe++l +iv+vhylevk
XP_011092424.1 115 RSYWLLEQDLMHIVFVHYLEVK 136
*******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.882 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 9.5E-81 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 9.4E-51 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.1E-4 | 434 | 521 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 2.0E-6 | 436 | 521 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 8.0E-16 | 436 | 522 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 19.89 | 619 | 740 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.51E-17 | 627 | 730 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.0E-16 | 629 | 732 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.80E-14 | 632 | 728 | No hit | No description |
| Pfam | PF12796 | 2.9E-6 | 636 | 695 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 7.4E-4 | 669 | 698 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.018 | 669 | 701 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2100 | 708 | 737 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.89E-7 | 841 | 892 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.84 | 841 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.675 | 843 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0064 | 844 | 862 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 4.6E-4 | 864 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.871 | 865 | 889 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.3E-5 | 867 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1021 aa Download sequence Send to blast |
MEESGSYNLG FRLDIKQILL EAQHRWLRPA EICEILRNYE KFHISPEAPN KPVSGSVFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDMLH CYYAHGEDNE NFQRRSYWLL 120 EQDLMHIVFV HYLEVKGNKT NLGGVRNNDR VVSNSENESS LSSSFRGTSP TSTLSSAYED 180 AESEGNHQAS SRFHSYPESP LTDDNHSAQS SSYNQLFNPG NQNVPALNYA SLLRGNRDGD 240 FGGDSLVCGA QETGDFALWQ EVLGNPTTGE IAYKPETGFS LPVQANRQAL NSLFEEKSLS 300 SDQGNDAGPF YSYPEQKGQS GENNLQMLLS DAEAGNVMNP NMENVMAAIG NENYSFLLKK 360 PLIGGLQTEE SLKKVDSFSR WMAKELGEAD GLDMQSSNGI SWSIIGNEYD SNMSAQLQVD 420 THTLNPSISQ DQLFSIIDFS PNWAYSNLDT KVLITGTFLK SEEELSNCRW SIMFGEVEVP 480 AQVLADGILC CRAPLHNPGL IPFYVTCSNR LACSEIREFE YRFGPDQNAD AVDVHGDSAI 540 LMHLYQRFET ILSLEPVGSP VSSAKNDLEK QSLVNKIISL MEENNPESKL TPNDDTSHLK 600 VIGELLLKKQ LRQIFYSWLL HRVTEDGKGL TVIDEGGQSV LHLAAALGFN WAFQPIIVSG 660 VSIDFRDVNG WTALHWAAFY GREDTVAALV SLGAAPGALT DPSAEYPRGR SPSHLASSRG 720 HKGISGFLAE TALTTHLSSL KVNDDCTKEV SGLKGILTVS ERSAVPTTEE DVPDTLSLKD 780 SLAAVCNATQ AAARIHQIFR VQSFQRKQLI EQDSDELLTP DEHAISLLAA KASRFNHSDG 840 VVNAAALQIQ KKYRGWKKRK EFLLIRQKIV KIQAHVRGHQ ARKKYKPIIW SVGILEKVIL 900 RWRRKGSGLR GFRSDAVQKG PDTPSLMPPQ EDDYDFLKEG RKQTEERMHK ELARVKSMAQ 960 YPEARAQYRR LLTAAQGFRE TKHDASDVIP DNMEDMIYPE EDLLDVASLL DDDTFMSLTF 1020 Q |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
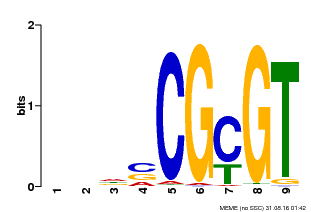
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011092424.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011092424.1 | 0.0 | calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A2G9H0E6 | 0.0 | A0A2G9H0E6_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.B00609.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



