 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011085456.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 822aa MW: 94918.7 Da PI: 7.0972 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.4 | 5.1e-28 | 65 | 152 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk+++dgkw+++++ +eHnHel p
XP_011085456.1 65 SFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESESG---SSRRPSVKKTDCKASMHVKRKRDGKWYIHEFIKEHNHELLP 152
5***************************************9999877...77888899******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.4E-25 | 65 | 152 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.2E-24 | 273 | 365 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.866 | 552 | 588 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.3E-6 | 553 | 585 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.3E-7 | 563 | 590 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 822 aa Download sequence Send to blast |
MVDYEDNVEC GEEGIGLNMV DVVDAGENRD GQVSSSHKGD ILGIEGDNFE PHDGIEFESH 60 EAAYSFYQEY AKSMGFTTSI KNSRRSKKTK EFIDAKFACS RYGVTPESES GSSRRPSVKK 120 TDCKASMHVK RKRDGKWYIH EFIKEHNHEL LPALAYHFRI HRNVKLAEKN NIDILHAVSE 180 RTRKMYVEMS RQSGGSFNSC LSKNEFDHQF DRGRYLALEE GDAQVMLEYF VQIQKENPGF 240 FYAIDLNEDQ RLRNLFWVDA KTRKDYISFS DAVFFDTSYI KINEKMPIAL FLGVNHHFQP 300 MLLGCALLAD ETKPTFVWLM KTWLRAMGGQ APTVIITDPD KQLKSAIEEV FPYSRHCFAL 360 WHILERIHET LAHVLKQHEN FMKKFNKCIF KSLTDEEFDM RWWKMVGRFE LQENEWVHSM 420 YVDRKKWVPT FMRDIFLAGL STYQRSESVN SFFDKYIHKK INLKEFVRQY GAILQNRYEE 480 EDMADFDTLH KQPALRSPSP WEKQMSTILT HAIFRKFQVE VLGVVGCHPK KEKEMGANVI 540 FRVDDCEKTE NFVVTWNEAK SEVSCSCLMF EYKGFLCRHS MIVLQICGLP SIPSRYILKR 600 WTKDAKSRQA VVDGNERIQT RVQRYNDLCK RAIELGEEGS LSEENYNIAC RALVEALKNC 660 VNVNNRTAAE CSNNSVGLRC AEEENQLLHV TKTNKKKNTN KKRKLQPEPE AVVIEASDSL 720 QQMENLSSEG ITLNGYYGSQ QHVHGLLNLM EPPHDAYYVS QQTMQGLGQL NSIASSHDGF 780 YGAQQSMPGL GHLDFRQPAF TYGMADEHNV RPAQLHGTAR HA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
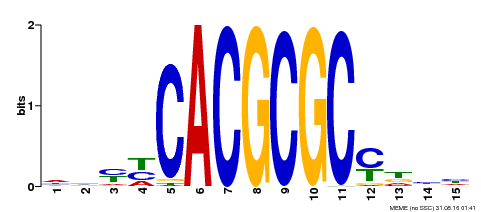
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011085456.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011085456.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011085457.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A022RW20 | 0.0 | A0A022RW20_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.H01263.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



