 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011082427.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 371aa MW: 41484.9 Da PI: 4.7256 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.2 | 3.3e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWT+eEde+l ++v+ G g+W++ ++ g+ R++k+c++rw +yl
XP_011082427.1 14 RGRWTAEEDEILRKYVEANGEGSWRSLPKNAGLLRCGKSCRLRWINYL 61
8*******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.9 | 1.8e-16 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+ +++E+e+++++++ lG++ W++Ia +++ gRt++++k++w+++l
XP_011082427.1 67 RGNISAQEEEIIINLHASLGNR-WSLIAGHLP-GRTDNEIKNYWNSHL 112
7899******************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-22 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.071 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.68E-28 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.4E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.7E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.86E-9 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.28 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.0E-25 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.4E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.6E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.39E-9 | 71 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009813 | Biological Process | flavonoid biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
MGRAPCCEKV GLKRGRWTAE EDEILRKYVE ANGEGSWRSL PKNAGLLRCG KSCRLRWINY 60 LRSDLKRGNI SAQEEEIIIN LHASLGNRWS LIAGHLPGRT DNEIKNYWNS HLSRKIQSFR 120 PTKFIPPAVA AAVRRTGRRT ARSVAKKNKN SELKGGGGAA NTPVPMPTTP TPEKEALSST 180 AEWQDKDNSC NVIPESSPLE ERLMESLGAA GTEESEIRNT TPVFSVTEEM EMENGDLCLD 240 DIMEDLNGIV GLDEERESPG KIEERESESC RNVDSGEKGS SGNGNHVVDD RGSWKSIGEW 300 NLMSSPGNSW EWDWCHDDDT LRGGNEEWGQ EENTLSWLWD GDTATGDCTD GDSEMDRRRE 360 KHNAMLSWLM S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-21 | 12 | 118 | 5 | 110 | B-MYB |
| 1gv2_A | 1e-21 | 12 | 116 | 2 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| 1h88_C | 4e-21 | 12 | 141 | 4 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 4e-21 | 12 | 141 | 4 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h8a_C | 2e-21 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| 1mse_C | 1e-21 | 12 | 116 | 2 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 1e-21 | 12 | 116 | 2 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
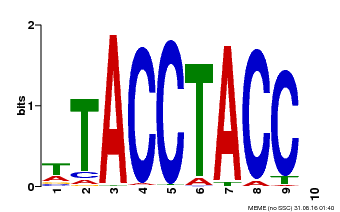
| Motif ID | Method | Source | Motif file |
| MP00627 | PBM | Transfer from PK06182.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011082427.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011082427.1 | 0.0 | myb-related protein Myb4 | ||||
| TrEMBL | A0A2G9GH61 | 1e-100 | A0A2G9GH61_9LAMI; Transcription factor, Myb superfamily | ||||
| STRING | EOX95226 | 3e-89 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G49330.1 | 1e-71 | myb domain protein 111 | ||||



