 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0224s0023.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 91146 Da PI: 6.3809 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.1 | 3e-18 | 9 | 67 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le++++++++ps +r++L k + +++ +q+kvWFqNrR +ek+
Sphfalx0224s0023.1.p 9 KYVRYTNEQVEALERVYHECPKPSSIRRQQLIKDCpilaNIEPKQIKVWFQNRRCREKQ 67
5678*****************************************************97 PP
| |||||||
| 2 | START | 162.3 | 3.7e-51 | 167 | 372 | 2 | 202 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT.. CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg.. 88
+aee+ +e+v+ka+ ++ W +++ +++g++ + +++ s++++g a+ra+g+v ++ v+e+l+d++ W +++ ++l + +g
Sphfalx0224s0023.1.p 167 IAEETVTEFVAKATGTAIDWIQLPGMKPGPDAIGIVSISHGCPGVAARACGLVGLEPN-KVAEILKDRPSWLPDCRCLDILGAFPTGng 254
7899******************************************************.8888888888******************** PP
EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE-- CS
START 89 galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlk 173
g+++l++++++a+++l+p Rdf ++Ry+ l++g++vi+++S++ +q p+ +s+vRae+lpSg+li+p+++g++ +++v+hvdl+
Sphfalx0224s0023.1.p 255 GTVELIYTQMYAPTTLAPpRDFCTLRYTTLLEDGSLVICERSLTGTQGGPTmprVQSFVRAEMLPSGYLIRPCDGGGCIIHIVDHVDLE 343
**************************************************999999********************************* PP
SSXXHHHHHHHHHHHHHHHHHHHHHHTXX CS
START 174 grlphwllrslvksglaegaktwvatlqr 202
++++++lr+l++s+++ ++kt+ ++++
Sphfalx0224s0023.1.p 344 PWSVPEVLRPLYESSAVLAQKTTIVAFRH 372
*********************98766655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.418 | 4 | 68 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.4E-15 | 6 | 72 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.15E-16 | 7 | 71 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.63E-16 | 9 | 69 | No hit | No description |
| Pfam | PF00046 | 8.9E-16 | 10 | 67 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.0E-18 | 11 | 67 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 9.79E-4 | 61 | 97 | No hit | No description |
| PROSITE profile | PS50848 | 25.559 | 157 | 358 | IPR002913 | START domain |
| CDD | cd08875 | 1.48E-76 | 161 | 375 | No hit | No description |
| SMART | SM00234 | 1.0E-41 | 166 | 376 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.0E-39 | 166 | 377 | No hit | No description |
| Pfam | PF01852 | 1.3E-48 | 167 | 372 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 9.7E-24 | 168 | 365 | IPR023393 | START-like domain |
| Pfam | PF08670 | 6.3E-44 | 694 | 836 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MQGMDASGKY VRYTNEQVEA LERVYHECPK PSSIRRQQLI KDCPILANIE PKQIKVWFQN 60 RRCREKQRKE ATRLVSVNAK LTALNKLLME ENERLAKHTS QLTVDNHFLR KQLQQSPPIP 120 GSKRPPDQAA VGSTDTSSDS VVTVGLPPQP TPQHSPRGTS PAGLLSIAEE TVTEFVAKAT 180 GTAIDWIQLP GMKPGPDAIG IVSISHGCPG VAARACGLVG LEPNKVAEIL KDRPSWLPDC 240 RCLDILGAFP TGNGGTVELI YTQMYAPTTL APPRDFCTLR YTTLLEDGSL VICERSLTGT 300 QGGPTMPRVQ SFVRAEMLPS GYLIRPCDGG GCIIHIVDHV DLEPWSVPEV LRPLYESSAV 360 LAQKTTIVAF RHLRRVAQEA SGAVGPRGGQ QPAVLRALSQ RLARGFNQAV NGFPDDGWAA 420 TVSDGMDDVS VVLNESPTAK GMGQVAADRL LCSLGGGILC AKASMLLQNV PPALLIRFLR 480 EHRSEWADCD IDADAAAAFR TSTRSGNAGH VQLPLPLAHS VEQEEFLEVV KLERHGGAPN 540 GIHTRDNFLL QFCSGVDENA VGACAQLVFA PVDVAVSDDV PLLPSGFRVI PLDCSLIDGL 600 GLSRTLDLAS TLEGGTEPSR ESGTGLNSSL RSVLTIAFQF AYEAHMREAC ANMARQYVRT 660 VVASVQRVAM ALAPSRLPLH LGAQQPPESP DVVTLTCHIL QSYRLHLGID LTRSDSGDEG 720 LFKAFWHHSD AIMCCAWKQG LPEFTFANQA GLDMLETTSG ALQDLSWEKT LDENGRKTAY 780 NDFTQVLQQG YVYLPAGVRI SSRGQPVTYE RALAWKVLAD DDTTRCIAFL FMNWSFFT* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
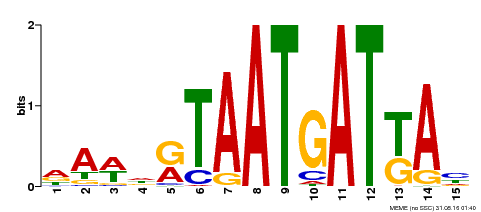
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KU530555 | 0.0 | KU530555.1 Sphagnum sp. SB-2016 C3HDZ1 mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024399219.1 | 0.0 | homeobox-leucine zipper protein HOX32-like isoform X2 | ||||
| Swissprot | A2XK30 | 0.0 | HOX32_ORYSI; Homeobox-leucine zipper protein HOX32 | ||||
| Swissprot | Q6AST1 | 0.0 | HOX32_ORYSJ; Homeobox-leucine zipper protein HOX32 | ||||
| TrEMBL | A0A0Y0G9Y6 | 0.0 | A0A0Y0G9Y6_9BRYO; C3HDZ1 (Fragment) | ||||
| STRING | PP1S15_11V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.2 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0224s0023.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



