 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0142s0001.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 607aa MW: 64465.2 Da PI: 4.3618 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 85.8 | 4.3e-27 | 293 | 347 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k++++WtpeLH+rFv+aveqL G ekA P++ilelm+v++Lt+++++SHLQkYR++
Sphfalx0142s0001.1.p 293 KAKVDWTPELHRRFVQAVEQL-GVEKAFPSRILELMGVQCLTRHNIASHLQKYRSH 347
6799*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.783 | 290 | 349 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.7E-27 | 291 | 350 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.96E-18 | 291 | 349 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.4E-25 | 293 | 347 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.9E-8 | 296 | 345 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 607 aa Download sequence Send to blast |
MKSCFVVAEE EEEQQQQAEE EKNNAPSPLL SQPPLFSVFE SEGPDCVVVD ALEVESLVTV 60 DDDGGSNAVV VAAAATAPEQ VAIHNDAPHV DDVQIAECFE FVSTSAGCGL DSEQEGGECE 120 DIDVSECLDF VQGRAAEDVS ECLDSSPPQA AAVAAAGGVV VGGGLLDGGE SISQCCMDFC 180 ESESFQDSFD VAQGYIDAAV EDAAASESLD SGGGGGDGVV EEVLRGRAVD IVDAAEVEAL 240 LFADVAKSTT TTTTTTAGEA VSGSDSGECS SGEKKEEERN HRTGATASGG KKKAKVDWTP 300 ELHRRFVQAV EQLGVEKAFP SRILELMGVQ CLTRHNIASH LQKYRSHRRH LAAREAEAAS 360 WSHNRRPYCN NTHHHPSSWS RSRRTEGSLP YLVPIHTNSP PPIQPRPSAA APHMVSSHAH 420 HQMQQPQPLM TPPPLKVWGY PTVDHSPVHM WQQSAMSPPS FWQAPDGSYW QHPAACYDAS 480 SHPPACYPHP MRVPMATMPS STGALLQAGF SSDQDSFFAD EALPTSMYLC CNSYESSDIG 540 RAAGAAACSN PPSESHLSKE ILDAAISEAL ANPWTPPPLG LKPPSMEGVI AELQRQGINT 600 VPPSTC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 4e-16 | 290 | 345 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
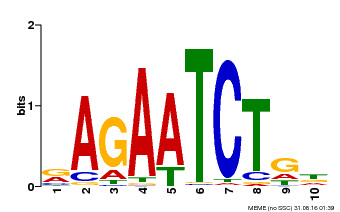
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024379514.1 | 1e-149 | transcription activator GLK1-like | ||||
| TrEMBL | Q5IFL7 | 1e-147 | Q5IFL7_PHYPA; Golden 2-like protein 1 | ||||
| STRING | PP1S311_30V6.1 | 1e-148 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.2 | 9e-32 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0142s0001.1.p |



