 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0111s0027.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 459aa MW: 49830.5 Da PI: 4.7181 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.4 | 1e-17 | 105 | 157 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++++ eq++ Le+ Fe ++++ e++ +LAk+lgL+ rqV vWFqNrRa++k
Sphfalx0111s0027.1.p 105 KRRLSFEQVRSLERNFELENKLEPEKKMQLAKELGLQPRQVAVWFQNRRARWK 157
4578889*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 120.6 | 7.8e-39 | 103 | 195 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrls eqv++LE++Fe e+kLepe+K++la+eLglqprqvavWFqnrRAR+ktkqlE+dye+L ++y++lk+e +++ e+e+L++
Sphfalx0111s0027.1.p 103 EKKRRLSFEQVRSLERNFELENKLEPEKKMQLAKELGLQPRQVAVWFQNRRARWKTKQLERDYEVLSQEYNRLKTEYDAVVLEKEKLES 191
69**************************************************************************************9 PP
HD-ZIP_I/II 90 elke 93
e+++
Sphfalx0111s0027.1.p 192 EIQH 195
9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.76E-19 | 89 | 161 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.508 | 99 | 159 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.6E-14 | 102 | 163 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 4.4E-15 | 104 | 157 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.61E-16 | 104 | 160 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-19 | 106 | 166 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.3E-5 | 130 | 139 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 134 | 157 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.3E-5 | 139 | 155 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.6E-14 | 159 | 199 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 459 aa Download sequence Send to blast |
MATTMRGFGA QNVMLMRNDS TPGSDSLVAM LASCTPMGLQ VPGHSSVGGL EDAVVGCGMK 60 RSFSYPSFVD HGSSSPGLGG EEALMGLDDG MDDCSPCSMS NVEKKRRLSF EQVRSLERNF 120 ELENKLEPEK KMQLAKELGL QPRQVAVWFQ NRRARWKTKQ LERDYEVLSQ EYNRLKTEYD 180 AVVLEKEKLE SEIQHLTGKS THKSSNMADQ SRDQYENVEP SAISGVKIKT TDHTGAKLSR 240 GADLQQVQVE VKSESMIRTD ASTEVVLKSR PADSQWKQQA STSPTVEIPA SILKAEKLEQ 300 QQTTSTSASN SSEILDTDSP RTIDNYKLQQ VSIITAAADQ QAGAGTVTSC QHLNNPGSAD 360 DQCFMDPDPA SCLLQVAAAG GGDDLMCAAG AADRVLSSPQ LQYHRSASVA ANIKLEVDAA 420 VAAGNCSNYQ AEYVDQSCNY FLSQLDDKAG ATPWWDWA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 151 | 159 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
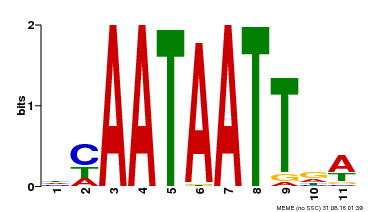
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 3e-42 | homeobox 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0111s0027.1.p |



