 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0079s0055.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 1032aa MW: 110253 Da PI: 6.0506 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.8 | 1.9e-16 | 69 | 113 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a k++G W++I +++g ++t+ q++s+ qk+
Sphfalx0079s0055.1.p 69 RERWTEEEHQRFLEALKLYGRA-WRRIEEHIG-TKTAVQIRSHAQKF 113
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.92E-16 | 63 | 119 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.755 | 64 | 118 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.1E-17 | 67 | 116 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.7E-13 | 68 | 116 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-13 | 69 | 112 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.7E-9 | 69 | 109 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.13E-9 | 71 | 114 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0009845 | Biological Process | seed germination | ||||
| GO:0009909 | Biological Process | regulation of flower development | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1032 aa Download sequence Send to blast |
MMIMTPSSAG DGEPHQAGGG GCGGEEGKLL QWSSSVAAAA VVGSSAAAAQ DDALENTKVR 60 KPYTITKQRE RWTEEEHQRF LEALKLYGRA WRRIEEHIGT KTAVQIRSHA QKFFSKVERD 120 HQAIACGIDT AGMVSQSLID IPPPRPKRKP SHPYPRKARG NSFSKHSDDN TTVVSSSSGL 180 ATEKSEEEEG FGKSSTMQED DTTTQVDEES SPPSLPKEKE DLGEDIKLPA VAPRQVLSEP 240 SSWVPFPPFH HYPPWGSCFQ NQVPHPNQHP AYGNLEKLAP FSGFVMPQEF MLPWHPSPMA 300 CTSSSVPQDL PAMAAPAVPT LWPAIGSETS AITTSDFSEI GSNTSSKGTE AVTSTAAMAA 360 AASVTIAAAS AWWALQNMAS PLSLTAMGGA LYNPVMAAIP SPLLFSEGQL LQPDCGGQNS 420 EPVAEDTPPL EQTLEDSSKE LLHSGHELVS SSAVQRPQRA VRVKEVKGRR GLAMMSSAGS 480 SDLCGSSSWG GHCLTPGSGG DSSREVGQRL ELLKSSGESI SRSPSNAGSS PTKLHKLKSQ 540 LSGLVQKELS SVHAKSSKDE GDEVGFEYWK HLRDESYSGF SMNPVTCSSP KLGVSGSPAF 600 QFLNSRGFSR FNSFPEECVN QDPYAKSSEE HQPALQTSSG SEASGSDGGI LAENASGEGG 660 GPSSNSSACG NLPEGNDPSG SSEGSSDVEV PSNVTSNCRE ENNGRSASEA EIVENEPEEA 720 QVSHFSHDAH LTTLRKDSEK HPQECAFRLL PRSLSLTSKK QIEAMGRGQI AFQALFAQDT 780 LPRTFTPPPR DSVAAHRTQA HDSGVVKQSL SFRLQSRISH SKASSFGSLP ADEDHPHHAI 840 DVAADQDSAF NAWQPSKPSE GSGAKIERCE SSSITMSSGA RVTKGAEEQQ SPQRTNNAKR 900 KERCGEGSAI NWLTLGTESS GLKQVKYNAE DQNFNSQREL LQKETFAPNI DFFPRQKKEK 960 SVASLKVHDT GSSAFVELSS VPLSGKTHMF PLSIATSGTA LSTISSTKSV KYSGLGFVPY 1020 QRASLEPSLQ L* |
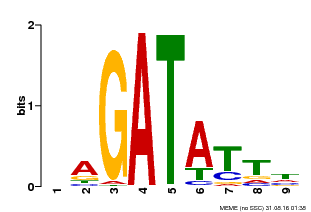
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G37260.1 | 2e-33 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0079s0055.1.p |



