 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0059s0014.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 766aa MW: 81816.3 Da PI: 6.8945 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.7 | 4.8e-32 | 119 | 172 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWt +LH++Fv+a+e+LGG+++AtPk +l+lm+vkgLt++hvkSHLQ+YR+
Sphfalx0059s0014.1.p 119 PRLRWTADLHHCFVHAIERLGGQDRATPKLVLQLMEVKGLTIAHVKSHLQMYRS 172
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 5.9E-31 | 115 | 173 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.711 | 115 | 175 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.36E-14 | 117 | 172 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.7E-23 | 119 | 173 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.0E-7 | 120 | 171 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 766 aa Download sequence Send to blast |
MQPKEERSNG SNEEEKVAGG NANSRKRSRH ASSSLSGGDE EEEYHHVVEQ SSESYNTAAT 60 GHGHGSSPGT AAAGRNHSCS HEEGGSYSSA AGGGGGGAGA AGSCRPGGTV RQYVRSKMPR 120 LRWTADLHHC FVHAIERLGG QDRATPKLVL QLMEVKGLTI AHVKSHLQMY RSMKIDENGH 180 SGGLPGAHDD PHQGLEFQDH AFDVGSCSFQ LMNPAASRSL FPAAGQTIDL HLGTSGNKSR 240 AAAGAFLQSE VTDFSPQRNI MISNFLQCHP PAPGVHVQHQ SDRLELDRRH LTMSLQRHDD 300 GRGGEWLQRP AAGVPTGIEI RSTATTAAGL IRDSWPHHGD QEKNEIVVNW ATRAGCQQAA 360 ALKAAVNGIT TSLQPGGAHS MQHLRTQQLV DEWSRKPSPA AVRLQADWDQ RQQVNHEAGG 420 GRAVAAHHES MQHQPLHLAM SQPATPAQGA GGPLKKQQRI TDHSREWERL CRGSSEHEGA 480 AVRPRMEWGS KITLDQGHRG GDGYASTSTA ASSIVSPFHL RQLLDSSCGS SNLQATGEVE 540 QRAAWSESQH GELLVAGGGT GQIISGWNMS EVQRKMALSA NPLQIRVGSL EKAEGDHGVE 600 SKSRFSAGGP PEYDPHPAAA IRRPWATTKM ETSLEAGRFW RAEEAAGAQS TTDHDPCGDQ 660 NLMQQQQDQQ LDTTTTTELS LSPFSSRGTK SCSPPAAAAA SCRMTKVTTL SLVPPGLTNE 720 TVESASADCD LSLQQQQSEM SDAVVNLQVP PKGITLDLTM SIGRP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 5e-16 | 120 | 174 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-16 | 120 | 174 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-16 | 120 | 174 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-16 | 120 | 174 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
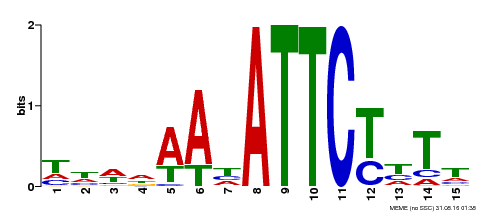
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 7e-31 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0059s0014.1.p |



