 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0047s0045.6.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 499aa MW: 51783.4 Da PI: 6.12 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36 | 1.2e-11 | 291 | 336 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P++ K +Ka++L + ++Y+k Lq
Sphfalx0047s0045.6.p 291 HSIAERLRRERIAERMKALQELVPNS-----NKTDKASMLDEIIDYVKFLQ 336
99***********************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 2.87E-13 | 284 | 340 | No hit | No description |
| SuperFamily | SSF47459 | 7.46E-17 | 284 | 345 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.453 | 286 | 335 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.3E-16 | 288 | 345 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.5E-9 | 291 | 336 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 7.2E-14 | 292 | 341 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 499 aa Download sequence Send to blast |
MDEFLEDMFA MPWDYNAGQG AMMMERAGTG AAMAGVEAAS NINHNSNIGG GSSMAAATQQ 60 QQKMFSMPLI QQQPRLLNNN NGSGALSGVN QAQLVSSQAS QLSSQQQQQG KEMRSEVCGM 120 EGVSVATQEP GSGSSTGLQM NNAAGLQQQQ QGGSLTGSAS VGSSGSEDSG GQLGRGDHSP 180 SPPTALTWQQ QQQPPYVGAV PPSLPMSLAQ AKVENILGRG EFNSHNAQML GKRFREDEEG 240 LLQENHSGIT GSGPTSAPSG QGFPGSSQQG LPTVGVRPRV RARRGQATDP HSIAERLRRE 300 RIAERMKALQ ELVPNSNKTD KASMLDEIID YVKFLQLQVK ILSMSRLGGA GAVASTASDH 360 PAEGSNPAAP TISRSLGTPT PVQDGIALTE RQVTRLMEDD MGSAMQYLQS KGLCLMPIAL 420 ATAISTTNSR GSHTATVGAG DRQKATAAGS NGGLAADGLV SEHSSKDNNR EVESTNGTTT 480 SMAIAKAPKG EGKPSDGP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 295 | 300 | RLRRER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
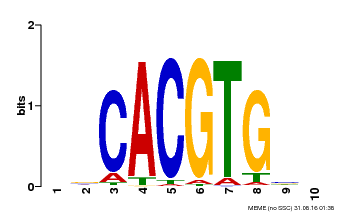
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024394844.1 | 1e-105 | transcription factor bHLH85-like isoform X2 | ||||
| TrEMBL | A0A2K1JI03 | 1e-104 | A0A2K1JI03_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S209_22V6.2 | 1e-105 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G24260.1 | 6e-47 | LJRHL1-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0047s0045.6.p |



