 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0046s0038.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1219aa MW: 133883 Da PI: 6.7534 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 170.4 | 2.6e-53 | 36 | 152 | 3 | 117 |
CG-1 3 kekkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahse 89
+ ++rwl++ e+++iL n+ k+ ++l ++ +p sgs++L++rk +ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+e
Sphfalx0046s0038.2.p 36 EAQSRWLRPLEVCEILSNYGKYGFKLnpVPPVKPCSGSMFLFDRKTLRYFRKDGHNWRKKKDGKTVREAHERLKTGSVDVLHCYYAHGE 124
459******************9998866899********************************************************** PP
CG-1 90 enptfqrrcywlLeeelekivlvhylev 117
+n+ fqrrcyw+L+ +le+ivlvhy+ev
Sphfalx0046s0038.2.p 125 DNSCFQRRCYWMLDPALEHIVLVHYREV 152
**************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.698 | 30 | 158 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.8E-72 | 33 | 153 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.2E-50 | 36 | 151 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.58E-18 | 625 | 711 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 3.3E-6 | 625 | 711 | IPR002909 | IPT domain |
| Gene3D | G3DSA:2.60.40.10 | 9.0E-5 | 625 | 712 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00204 | 2.79E-16 | 779 | 918 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.5E-19 | 807 | 923 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.91E-18 | 810 | 921 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.794 | 826 | 922 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.7E-7 | 831 | 920 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.831 | 859 | 891 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0017 | 859 | 888 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3000 | 898 | 929 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.19E-7 | 1028 | 1095 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.29 | 1044 | 1066 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.553 | 1045 | 1074 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0083 | 1048 | 1065 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.17 | 1067 | 1089 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.755 | 1068 | 1092 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.016 | 1070 | 1089 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 24 | 1143 | 1165 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.968 | 1144 | 1173 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1219 aa Download sequence Send to blast |
MEGVSENRAR LRVVTAAAQH HQQQQAEVDI RQILAEAQSR WLRPLEVCEI LSNYGKYGFK 60 LNPVPPVKPC SGSMFLFDRK TLRYFRKDGH NWRKKKDGKT VREAHERLKT GSVDVLHCYY 120 AHGEDNSCFQ RRCYWMLDPA LEHIVLVHYR EVTESSRFST MAADTTTTTH SSSSPSPSVT 180 VAGSPDVSGV GPSEQEHEDG DVVESDEVED SSSSSLLDFG LATPIQQQQP SSLLEQFAPP 240 PPPPQPSPPP GLPSGLDTKV FHSWGAPPPP PLNSSASPAS SAGLVGIEKL LNSSSSSSSS 300 SGVAPSSTSF FSQLNRQQQQ PSLLRMDYLG RSQQRESTGG LPDLLNNNNN NNPDPFLGGE 360 LLPRIQPSST TFDITGWSEL LENSQRNHAA SKNINQNIAA SGMMVGEAVY VKQESMNGSA 420 WDMLEQCITP STQRNIVESI MANSSSVEGD RGGLEVAANG QDLTFGSLIP SPKGILEALS 480 PRGLGREPQT PLEASLRAVT EEKAIRAAAE AQQRRFAGGS TRSDPQNADA LWDAKAAEVA 540 FSKLESFKNS ELGNLTKFDS FGRWMINELG RDGQNFLPSG PSQSGWSGMG PLEDQGVYDM 600 YSFSQQMQVE AGLSTPVSQD QRYSIVDFSP EWAYPAGDAK VIITGMFLRE YKSSSEDYKW 660 CCMFGELEVP AEVIGPGVLR CKAPPHALGR VSFYVTCGDR QAHSEIREFE YCSAASTAVG 720 LAATKSGLKT DEEMMLKVRL ARMLLAESST QDSGMEALPG TAEILNSVYS DDEWSYLEHL 780 AKSSGDLQAF NFQEQLLQML LKKHMQKWLL SKVQEDGKGP SILDEQGQGV FHMAAALGYD 840 WAVAPLVAAG IPINFRDVRG WTALHWAASH SREEVVIALL KARADPGTVT DPTRAFPAGQ 900 TPADLAAANG HGGIAGFLAE ANLIGRLSSM TISESPMDIA MSHIAGEKAV AKLSRRESIR 960 RSMNGGEDEL SVVESLQAVR NAAQAAALIN AAFQRHSFQK QEEDSLASIE ADEFGMTPRQ 1020 LRGLMAARTA QKFQNAFRGH HEKKQQLAAT RIQRKYRGWK QRRTFLDLRQ RVVKLQAQVR 1080 GNMVRKRLKK LLWSVGILEK AVLRWRRKGA GLRGFKSGEL SVPDKEEDDE ELLKDGRKKN 1140 EAALEKAVTR VQSMVRSRQA RAQYLRLRDG SQQQGQFSQP STPQDFTEDQ SISFQQSQHQ 1200 GRSLRSQSPL GFSDSMII* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 9e-14 | 625 | 718 | 9 | 95 | calmodulin binding transcription activator 1 |
| 2cxk_B | 9e-14 | 625 | 718 | 9 | 95 | calmodulin binding transcription activator 1 |
| 2cxk_C | 9e-14 | 625 | 718 | 9 | 95 | calmodulin binding transcription activator 1 |
| 2cxk_D | 9e-14 | 625 | 718 | 9 | 95 | calmodulin binding transcription activator 1 |
| 2cxk_E | 9e-14 | 625 | 718 | 9 | 95 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
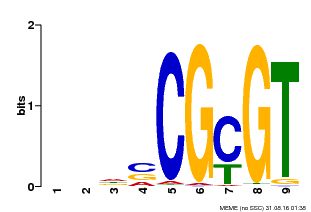
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024381994.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| Refseq | XP_024381996.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| Refseq | XP_024381997.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| TrEMBL | A0A2K1K5Y1 | 0.0 | A0A2K1K5Y1_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S188_92V6.2 | 0.0 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 1e-121 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0046s0038.2.p |



