 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0023s0153.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 559aa MW: 59601.3 Da PI: 5.1884 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 34.7 | 3.1e-11 | 482 | 527 | 6 | 55 |
HHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 6 nerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+++Er RR +i +++ +L++l+P+ +k + +++L +AveY+k+Lq
Sphfalx0023s0153.1.p 482 SIAERVRRTKISERMKRLQDLVPNM----DKQTNMSDMLDEAVEYVKQLQ 527
689*********************7....5779999*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 14.488 | 476 | 526 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 4.97E-16 | 479 | 542 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.67E-12 | 480 | 531 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 7.5E-17 | 481 | 542 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.1E-8 | 482 | 527 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.8E-13 | 482 | 532 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 559 aa Download sequence Send to blast |
MAGMSQVDEE LSAMLQSDLH QSPAQTTLAS VMSSPAGMVL GPGSEHMDSS TFSSHYSSSL 60 EGGGLISSSS SGGSHGRQQL LVPPTVRGAT NLSSVEVAAS SHVLSLAAAA GPVQCSRPME 120 QQQLLWPFIA ADQKLLELQD PATKNQDFMK KAVNSQVVNF STSLLDATSM SLQDLNPPPP 180 QFSSRGGLQR YHSAPSSLLR SLAADCNNED VLGFSSHQVS SPLGNNTGNI RDDTRGRSLI 240 PPFFSGAGNN NNNLTSITEC VVQPMETDKM LNDSTLHQYD RFLSPATKSN KRASLVTTTS 300 MLAGKSDSKP ADLFGNQPQL GNDNLLRQSS LPADWFAALQ GSIDDTDPSS QMPSMTSPDE 360 SLSGRTSEDT AGVGASGKSS IYSRDDSFLV IDTTSGGLWD SPTSPVGHIG GKRLRGILGD 420 ATIDTGEGLQ RDRLGGPGTL IRHTSLPANF NRPTIPVLDD LADISVPCQT RAKRGCATHP 480 RSIAERVRRT KISERMKRLQ DLVPNMDKQT NMSDMLDEAV EYVKQLQQQV QDLSDTVAQL 540 RDLQAKGLKT EDNGSSEV* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00181 | DAP | Transfer from AT1G35460 | Download |
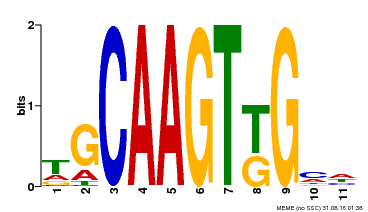
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G35460.1 | 2e-35 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0023s0153.1.p |



