 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0022s0100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 476aa MW: 51387.9 Da PI: 4.7946 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.5 | 4.8e-18 | 121 | 174 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++t +q++ Le+ Fe ++++ e++ +LAk+lgL+ rqV vWFqNrRa++k
Sphfalx0022s0100.1.p 121 KKRRLTFDQVRSLERNFELENKLEPERKMQLAKELGLQPRQVAVWFQNRRARWK 174
55678999*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 121.8 | 3.4e-39 | 120 | 211 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrl+ +qv++LE++Fe e+kLeperK++la+eLglqprqvavWFqnrRAR+ktkqlE+dye L + y++lk+e e++++e++eL++
Sphfalx0022s0100.1.p 120 EKKRRLTFDQVRSLERNFELENKLEPERKMQLAKELGLQPRQVAVWFQNRRARWKTKQLERDYEILTQDYNRLKSELESVRDEKQELQA 208
69**************************************************************************************9 PP
HD-ZIP_I/II 90 elk 92
++k
Sphfalx0022s0100.1.p 209 KVK 211
987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 8.6E-20 | 102 | 177 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.3E-19 | 113 | 178 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.443 | 116 | 176 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 7.0E-15 | 119 | 180 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.05E-15 | 121 | 177 | No hit | No description |
| Pfam | PF00046 | 1.9E-15 | 121 | 174 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 1.1E-5 | 147 | 156 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 151 | 174 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.1E-5 | 156 | 172 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.7E-13 | 176 | 216 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 476 aa Download sequence Send to blast |
MMQQRQANNN LVRGFATGPS DMMLMSNRSE NSSTDTLAAM LASCTPAGAL QASRSSGSLE 60 DAVMGCGSGG GGGQKRPSSF YPPPPPTTST TFDASLLLDA AGMDAADDGG DEYCGLHNAE 120 KKRRLTFDQV RSLERNFELE NKLEPERKMQ LAKELGLQPR QVAVWFQNRR ARWKTKQLER 180 DYEILTQDYN RLKSELESVR DEKQELQAKV KWLTEKSTTQ PEEESAAGCQ AAPIGSYVQQ 240 QQSSSKLVET TATTLNHQPG ATSNESPSEQ LTLPEKPSGA VVEAHVFSRS SSCRDMVDDQ 300 PTRLTAFINP VSSSAEVMPS SSAATRTRKR KDGSPSSAYS TSSEILDADS PRTIDSGLNS 360 HTIMSSTVTS STVAASSACP GQMYVDTAAA TTTTIVNVPS NSCLNVDLVM GQDLVCSRTF 420 SDPQTYNQRS TTSAVKLEGA INGAAMVFQP ADQDSCNYLL PQVDEHGALP WWDWA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 168 | 176 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
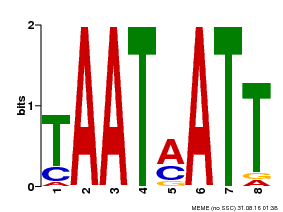
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 5e-38 | homeobox protein 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0022s0100.1.p |



