 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0012s0198.6.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 758aa MW: 81601 Da PI: 6.33 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 89.5 | 3e-28 | 243 | 296 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Sphfalx0012s0198.6.p 243 KPRVVWSVELHQQFVSAVHQL-GIDKAVPKRILELMSVHGLTRENVASHLQKYRL 296
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 80.8 | 4.6e-27 | 57 | 165 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedal 87
vl+vdD+p+ +++l+++l + y +v+++ +++alell+e + +Dl++ D+ mp+mdG++ll+ e++lp+i+++a ge + +
Sphfalx0012s0198.6.p 57 VLVVDDDPICLMILERMLHQCSY-QVTTCGRATKALELLREDKdkFDLVISDVYMPDMDGFKLLELVGL-EMDLPVIMMSADGETSAVM 143
89*********************.***************888778*******************87754.558**************** PP
HHHHTTESEEEESS--HHHHHH CS
Response_reg 88 ealkaGakdflsKpfdpeelvk 109
+ + Ga d+l Kp+ eel++
Sphfalx0012s0198.6.p 144 KGITHGACDYLLKPVRLEELRN 165
********************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF52172 | 1.85E-37 | 54 | 176 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 4.8E-44 | 54 | 189 | No hit | No description |
| SMART | SM00448 | 1.4E-32 | 55 | 167 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 45.545 | 56 | 171 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 4.8E-24 | 57 | 166 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 2.10E-29 | 58 | 171 | No hit | No description |
| PROSITE profile | PS51294 | 11.526 | 240 | 299 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.48E-19 | 240 | 300 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-29 | 242 | 301 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.6E-24 | 243 | 296 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.8E-7 | 245 | 295 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009736 | Biological Process | cytokinin-activated signaling pathway | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 758 aa Download sequence Send to blast |
MASVQHPLSH QLLPGHEQQR QQQQYGGAST GTHNMLQAHN YHKDVDSVDE FPAGLRVLVV 60 DDDPICLMIL ERMLHQCSYQ VTTCGRATKA LELLREDKDK FDLVISDVYM PDMDGFKLLE 120 LVGLEMDLPV IMMSADGETS AVMKGITHGA CDYLLKPVRL EELRNIWQHV LRKKRTGAKD 180 VEHSTSHEEG ERRKFGGGED VEYTSSATDT TDGNCKLMKR RKEMKEDDED TEQENDDPST 240 LKKPRVVWSV ELHQQFVSAV HQLGIDKAVP KRILELMSVH GLTRENVASH LQKYRLYLRR 300 LSGVTSQQNG MNVMFGGSET AFGHVSSLDS IGNLRTLAAT GQLSPQTLAG VHASRIGREG 360 SPNCMGLPSS LDPSMSAQIA APPRPHMNGG LFGTQSCSVQ SLAGGVDVDP ILQLHNLSAM 420 GRLHQLDDIP GLRSLQHQLV GTSAGNCNTL GLNPVGGPSV NLVGRPSSEA LLMQLFQQRA 480 QQQGGGLSVH LPQPSAVLGT PCLLSTGVNL GKAGSLADMG REFGSSLGAA VASIPSSLGS 540 LVQPPDVGVQ TSNLVGSLGG SFNGSATVLS PTIGGAPGPS KMNDMPRPQH SMKTSLNPLG 600 QASHMHGQVL SQSSRPTWLG SRGMVFAEQG QSIESGLSPW FPSQNQAHVG RFAENLQQLG 660 QPDYTGQGMN FPVSLGLDIK PSIDGSQGQS GQPATCSEQK LLKDERASDN IIITKVDGGL 720 VPHDQSEDLL TFFFKQSHDG ISFPDSDGYR VDNFYVK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 6e-21 | 242 | 302 | 4 | 64 | ARR10-B |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
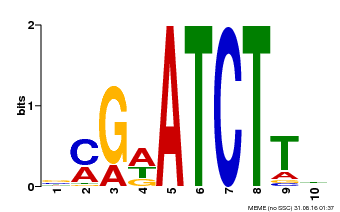
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024363770.1 | 1e-170 | two-component response regulator ARR12-like | ||||
| TrEMBL | A0A2K1IGK7 | 1e-168 | A0A2K1IGK7_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S362_27V6.1 | 1e-167 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-115 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0012s0198.6.p |



