 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0012s0197.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 477aa MW: 51957.2 Da PI: 4.5418 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 47.7 | 3.9e-15 | 65 | 114 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++ GVr+++ +grWvAeI+d + ++ r +lg+f ae+Aa+a+++a+ l+g
Sphfalx0012s0197.1.p 65 RFLGVRQRP-SGRWVAEIKDTT---QKIRLWLGTFNCAEDAARAYDEAAWMLRG 114
577******.7*********32...35*********************976665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.94E-27 | 64 | 121 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 8.2E-26 | 65 | 121 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.6E-32 | 65 | 128 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.019 | 65 | 122 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.3E-6 | 66 | 77 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 9.15E-18 | 66 | 122 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.2E-10 | 68 | 114 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.3E-6 | 88 | 104 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0035865 | Biological Process | cellular response to potassium ion | ||||
| GO:0048528 | Biological Process | post-embryonic root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 477 aa Download sequence Send to blast |
MEQQQQWQQV TNCGSVLELK HAASDSVKGR KEERESVAAR GASSLKKNSS YSMANCRGRS 60 RSNKRFLGVR QRPSGRWVAE IKDTTQKIRL WLGTFNCAED AARAYDEAAW MLRGSNTRTN 120 FLLPPPQAPS SSSANNFSLL PSKSARLLHR RRSSAAAAKL ESTEEADAAA AHVAAISAAA 180 AAAAASTKEV DSSETDDSLT SAAFEVVPTT PNALKVAITS GASETEDSLV AGAERELSYR 240 SSSSSNTSSA SDMISDSTWS YSSTEESLTI FPEMDHKEWL KAFSATEEQQ QYLEEGGAAM 300 MSDSGCSSSY RSMEESLMIS FPEMPGNRVW LEPIRRERQY LETKPLQEDD DETSTGFVAT 360 AAEDSSDELQ YYTTSFDFSA TDLEPLEKVP DTSSLEDPAL SDDTMLLNEH FLRMKEDTIL 420 GAGAWNQKLL EYAEEDKILG AGAWNQSCLK YAKEDNILGA GALNHKVSST SKSCIL* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00521 | DAP | Transfer from AT5G19790 | Download |
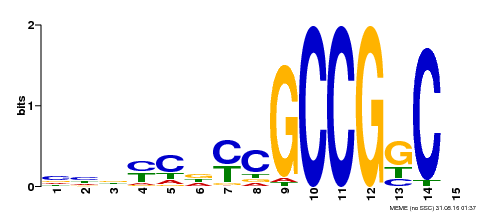
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G19790.1 | 2e-19 | related to AP2 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0012s0197.1.p |



