 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0008s0122.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 549aa MW: 59024.5 Da PI: 5.063 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 54.9 | 1.5e-17 | 141 | 193 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++++ +q++ Le+ Fe ++++ e++ +LAk+lgL+ rqV vWFqNrRa++k
Sphfalx0008s0122.1.p 141 KRRLNFDQVKSLEKNFELENKLEPERKIQLAKELGLQPRQVAVWFQNRRARWK 193
4466779*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 115.3 | 3.7e-37 | 139 | 230 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrl+ +qvk+LE++Fe e+kLeperK +la+eLglqprqvavWFqnrRAR+ktkqlE+dy++L y++lk++ e++ +e++eL++
Sphfalx0008s0122.1.p 139 EKKRRLNFDQVKSLEKNFELENKLEPERKIQLAKELGLQPRQVAVWFQNRRARWKTKQLERDYQVLSLDYNRLKNQLEAVLQEKQELQA 227
69**************************************************************************************9 PP
HD-ZIP_I/II 90 elk 92
+++
Sphfalx0008s0122.1.p 228 KVQ 230
987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.36E-19 | 127 | 197 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.167 | 135 | 195 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.3E-14 | 138 | 199 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.07E-16 | 140 | 196 | No hit | No description |
| Pfam | PF00046 | 6.8E-15 | 141 | 193 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-18 | 143 | 202 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 2.0E-5 | 166 | 175 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 170 | 193 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 2.0E-5 | 175 | 191 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 5.6E-11 | 195 | 234 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 549 aa Download sequence Send to blast |
MGAMDYNLQC PVPGTAVCFQ PADQMMANTV AGSRSSFGAI GTPNMQQMLM PSDQSSSTDT 60 LVAMLASYSP AASAALQASL SSGRLEDIAL VGCGQMRPSF LQYPASAYED SYLDEDQAPA 120 GDDDSAGDVS AAGHNNYVEK KRRLNFDQVK SLEKNFELEN KLEPERKIQL AKELGLQPRQ 180 VAVWFQNRRA RWKTKQLERD YQVLSLDYNR LKNQLEAVLQ EKQELQAKVQ CLTTAKAAGQ 240 EPSAKEERST GRVEEAAATT GLVDHHVLQS CKLEGLAMAP DVQTALRRSN SKSCNIAVDS 300 SQLNVKPDDQ YNSSKDADVD QVLISTTRDH VKKNRLKRAA GSRTNFAAAA VADADQAATT 360 AAPAVTPSPT LARKTRNMND GSPSSASTYS SDILDAEDSP HGTINSCTEL VSRNLTSASS 420 ISMRAAADSQ IRVQEHHPSH GVLDRYYQVT DQNLHYAGRV MNTFPHYEAE AEAALYNNVL 480 VGPRCTSTSV VKLEDVQVAA ASSAVFQEAA AADIAACNYL LAAAQVQEEH GGGGGGGGGA 540 LEPWWDWP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 187 | 195 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
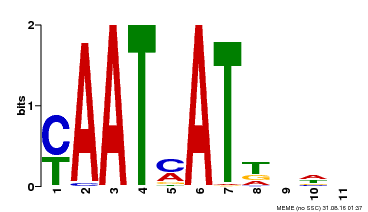
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 8e-26 | homeobox 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0008s0122.1.p |



