 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sphfalx0005s0290.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Sphagnophytina; Sphagnopsida; Sphagnales; Sphagnaceae; Sphagnum
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 615aa MW: 65085.7 Da PI: 7.8138 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 38.2 | 2.8e-12 | 132 | 172 | 1 | 42 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkeek 42
k++k+CnCk+s+Clk+YCeCfa+g++C+ C+C +C+N+ e+
Sphfalx0005s0290.1.p 132 KKCKQCNCKNSRCLKLYCECFASGTYCEG-CNCVNCCNNVEN 172
589*************************9.********9875 PP
| |||||||
| 2 | TCR | 51.4 | 2.1e-16 | 216 | 255 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
k++kgC+Ckks ClkkYCeCf+a+ Cs++CkC dCkN e
Sphfalx0005s0290.1.p 216 KHNKGCHCKKSGCLKKYCECFQANILCSDNCKCVDCKNFE 255
589***********************************65 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 4.4E-13 | 132 | 172 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 36.321 | 133 | 256 | IPR005172 | CRC domain |
| Pfam | PF03638 | 1.7E-8 | 135 | 169 | IPR005172 | CRC domain |
| SMART | SM01114 | 2.0E-20 | 216 | 257 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 6.3E-13 | 219 | 254 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 615 aa Download sequence Send to blast |
MAEEPPEQIA GVKTDAPGSP GGIVAAASSN GAHQVQSMAF APTSPAVSRT VNRPVDLDFP 60 QPVDTVSGPM QGVQGDHATR KAVPRQLDFT SMYGGPAGSA EGPVRTQPPV KHGTPRAERS 120 RPAFDSKDGT PKKCKQCNCK NSRCLKLYCE CFASGTYCEG CNCVNCCNNV ENESVRQEAV 180 EATLERNPNA FRPKIFSSPS LRDLREDVGE QPLIGKHNKG CHCKKSGCLK KYCECFQANI 240 LCSDNCKCVD CKNFEGSEER RALFHGDHNM GLNFMVQMAA NGSSQTIPGY LSPSPPMKKR 300 RTHELVFAGP GLKEQGPPGR TPSLPVQGHL AATVSTAVNT PITSMSTIAG GSASAAPVKV 360 VHRSLLAGVV QLEAIQELCK LLVIVSAEAQ KDFLAKHSLP CITAQQATET GGSKSSKEPS 420 PGIGREEEVP KQKSDERSDP HESEKASPAR MVSAVGDDVE VDTATQQRAM SPGTLALMCD 480 EKDPLFTAPP SPSGGLCSTE FSNTTPSHPG QLYAEQERAI LLEYRECLRR IIAVGKRRAS 540 QFATEVAQAE MYAATSLQQR GQPGSDGAQH PRPVAAPSAP VIVGSTGLCT IGMKPAKLGL 600 PLQTAAAPTS SVSQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 2e-33 | 135 | 265 | 12 | 132 | Protein lin-54 homolog |
| 5fd3_B | 2e-33 | 135 | 265 | 12 | 132 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 298 | 302 | KRRTH |
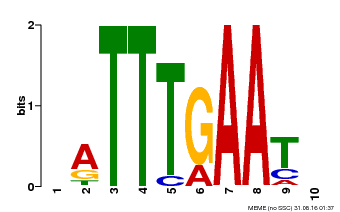
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00045 | PBM | Transfer from AT4G29000 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024361559.1 | 0.0 | protein tesmin/TSO1-like CXC 5 | ||||
| TrEMBL | A0A2K1IIF0 | 0.0 | A0A2K1IIF0_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S10_254V6.2 | 0.0 | (Physcomitrella patens) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29000.1 | 1e-107 | Tesmin/TSO1-like CXC domain-containing protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sphfalx0005s0290.1.p |



