 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.004G351700.1.p | ||||||||
| Common Name | Sb04g037850, SORBIDRAFT_04g037850 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 392aa MW: 40479.4 Da PI: 7.6063 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.9 | 0.00068 | 172 | 194 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y+C+ C+k+F++ L H +H
Sobic.004G351700.1.p 172 YECKTCNKCFPTFQALGGHRASH 194
99**********99999998887 PP
| |||||||
| 2 | zf-C2H2 | 11.5 | 0.00093 | 280 | 302 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C++Cg F + L H+r+H
Sobic.004G351700.1.p 280 HECSICGAEFASGQALGGHMRRH 302
78********************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF13912 | 2.3E-11 | 171 | 196 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.7E-8 | 171 | 194 | No hit | No description |
| PROSITE profile | PS50157 | 10.097 | 172 | 199 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.094 | 172 | 194 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 174 | 194 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.7E-8 | 275 | 302 | No hit | No description |
| Pfam | PF13912 | 1.8E-12 | 279 | 303 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.64 | 280 | 307 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.016 | 280 | 302 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 282 | 302 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 392 aa Download sequence Send to blast |
MQTSSELQLL VAQRNKTTNT NTKTAALTTL LMTTPTPTPT SSSTDDHSTA DLAQVDDDSH 60 GHHQPLAAAV KRKQRTKRPR HHPPAAASSC ASASVSSSES TTTEEEDMAH CLILLAQGAS 120 VVDSKPSPPV VLPSSTAPHQ SQAPPRAERY TSRKYTEAAA TPDGVRAGFY VYECKTCNKC 180 FPTFQALGGH RASHKKPRLA ATADGDIAAA VNVDGTTTTG TMATASPSAP PPLVVQPRPL 240 QTAAIDAAAA ATAGAFPDVT ITTALSLSSV ATASGKLRVH ECSICGAEFA SGQALGGHMR 300 RHRPLNAPDR AVTVTTAIVA ADTTGNSNSK KESSAGINLE LDLNLPAPSD EEAVVSRLPP 360 PPAVMLGLGQ FNDGKKAGLM LTASALVDCH Y* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.8870 | 0.0 | shoot | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00487 | DAP | Transfer from AT5G04390 | Download |
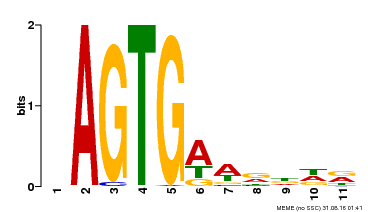
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.004G351700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP003989 | 7e-60 | AP003989.4 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OJ1063_D06. | |||
| GenBank | AP004026 | 7e-60 | AP004026.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OJ1136_C04. | |||
| GenBank | AP014958 | 7e-60 | AP014958.1 Oryza sativa Japonica Group DNA, chromosome 2, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012610 | 7e-60 | CP012610.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 2 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002454807.1 | 0.0 | zinc finger protein ZAT5 | ||||
| TrEMBL | C5XWL0 | 0.0 | C5XWL0_SORBI; Uncharacterized protein | ||||
| STRING | Sb04g037850.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2013 | 32 | 85 | Representative plant | OGRP2350 | 10 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G04390.1 | 7e-22 | C2H2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.004G351700.1.p |
| Entrez Gene | 8059712 |



