 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.003G411900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 811aa MW: 90774.6 Da PI: 6.4214 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.3 | 3e-23 | 128 | 229 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvF 82
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d++g +W++++i+r++++r++l++GW+ Fv++++L +gD ++F
Sobic.003G411900.1.p 128 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHGMEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIF 214
99*****************************7555544..58*********************************************** PP
EE-SSSEE..EEEEE-S CS
B3 83 kldgrsefelvvkvfrk 99
+ +++el+v+v+r+
Sobic.003G411900.1.p 215 L--RGENGELRVGVRRA 229
*..449999*****996 PP
| |||||||
| 2 | Auxin_resp | 112.5 | 4e-37 | 254 | 335 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++s+GmRf+m+fe+e+++e+r++Gt+vg ++ldp Wp+S Wr Lk
Sobic.003G411900.1.p 254 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSIGMRFRMRFEGEEAPEQRFTGTIVGCENLDP-LWPDSSWRYLK 335
89*********************************************************************.9*******996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.9E-41 | 117 | 243 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.29E-39 | 123 | 257 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.89E-21 | 126 | 228 | No hit | No description |
| Pfam | PF02362 | 4.0E-21 | 128 | 229 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.928 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.8E-23 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.4E-33 | 254 | 335 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 4.6E-6 | 677 | 734 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 4.81E-13 | 692 | 775 | No hit | No description |
| PROSITE profile | PS51745 | 24.764 | 698 | 791 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 7.0E-8 | 743 | 783 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 811 aa Download sequence Send to blast |
MPPAAAMALP SQAPSNSGDP LYPELWRACA GPLVTVPRVG DLVFYFPQGH IEQVEASMNQ 60 VAGNQMRLYD LPSKLLCRVL NVELKAETDT DEVYAQIMLM PEPEQTDVAA EKASSASAAS 120 PRPAVRSFCK TLTASDTSTH GGFSVLRRHA DECLPPLDMT QSPPTQELVA KDLHGMEWRF 180 RHIFRGQPRR HLLQSGWSVF VSSKRLVAGD AFIFLRGENG ELRVGVRRAM RQLSNVPSSV 240 ISSQSMHLGV LATAWHAINT KSMFTVYYKP RTSPSEFIIP YDQYMESVKN NYSIGMRFRM 300 RFEGEEAPEQ RFTGTIVGCE NLDPLWPDSS WRYLKVRWDE PSTIPRPDRV SPWKIEPASS 360 PPVNPLPLSS RVKRPRQNAP PPSPEASVLT KESAAKIDID SAQTQHQNSV LQGQEQMTLR 420 NNLTESNDSD STVQKPMMWS PSPNGKAHTN FQQRPAMDSW MPMGRRETDF KDSRSAFKDA 480 RTASQSFRDT QGFFMQAYDD NHHRLSFNNQ FQDQGSAHRF ADPYFYMAQQ PSLTVESSTR 540 TQTANNDLRF WGDQNSIYGN PGDQQQQGFS FGQNPSSWLN QPFPQVEQPR VVRPHATVAP 600 FDLEKTREGS GFKIFGFQVD TTSPSPAQLS SPLCAIREHV VQTRPSAPVN ELQPVQNECL 660 PEGSVSTAGT ATENEKNIQQ AQPSSKDIQS KSQGASTRSC TKVHKQGVAL GRSVDLSKFT 720 DYDELKAELD KMFEFNGELV SANRSWQIVY TDNEGDMMLV GDDPWEEFCS IVRKIYIYTK 780 EEVQKMNSKS SAPRKEEPLT VGEGCAATNE * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-165 | 19 | 360 | 17 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-165 | 19 | 360 | 17 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-165 | 19 | 360 | 17 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-165 | 19 | 360 | 17 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-165 | 19 | 360 | 17 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.3872 | 0.0 | callus| embryo| leaf| ovary| panicle | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
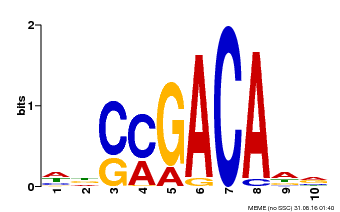
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.003G411900.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT067605 | 0.0 | BT067605.1 Zea mays full-length cDNA clone ZM_BFc0099C24 mRNA, complete cds. | |||
| GenBank | KJ726815 | 0.0 | KJ726815.1 Zea mays clone pUT3354 ARF transcription factor (ARF25) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021311466.1 | 0.0 | auxin response factor 4 | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A1B6Q7Z0 | 0.0 | A0A1B6Q7Z0_SORBI; Auxin response factor | ||||
| STRING | Si000340m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 | Representative plant | OGRP550 | 15 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.003G411900.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



