 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.003G144200.1.p | ||||||||
| Common Name | Sb03g012480, SORBIDRAFT_03g012480 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 295aa MW: 32458.5 Da PI: 5.681 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.7 | 2.2e-09 | 228 | 261 | 22 | 55 |
SSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 22 nrypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++e++ +LA+ +gL+ +q+ +WF N+R ++
Sobic.003G144200.1.p 228 WPYPTEEDKVRLAAMTGLDPKQINNWFINQRKRH 261
69*****************************885 PP
| |||||||
| 2 | ELK | 33.2 | 1.1e-11 | 181 | 202 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsg+L+ L++EF+
Sobic.003G144200.1.p 181 ELKEMLLKKYSGCLSRLRSEFL 202
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 5.3E-21 | 36 | 80 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 4.9E-20 | 38 | 78 | IPR005540 | KNOX1 |
| SMART | SM01256 | 8.7E-27 | 85 | 136 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.6E-23 | 89 | 135 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 10.423 | 181 | 201 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.3E-8 | 181 | 202 | IPR005539 | ELK domain |
| SMART | SM01188 | 3.6E-6 | 181 | 202 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.633 | 201 | 264 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 8.55E-20 | 203 | 272 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 4.4E-14 | 203 | 268 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-27 | 206 | 266 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.14E-13 | 213 | 265 | No hit | No description |
| Pfam | PF05920 | 4.8E-17 | 221 | 260 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 239 | 262 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010073 | Biological Process | meristem maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MEDLYSIHPG ISRVGGAASE ASVAGVGGPS PSDLTELMKA QIASHPRYPS LLSAYIECRK 60 VGAPPQVASL LEEVSRDRER RPGAGAGEIG VDPELDEFMD SYCRVLVRYK EELSRPFDEA 120 ASFLSSIQAQ LSNLCSAGSS PAATATHSDD MMGSSEDEQC SGDTDVPDIG QEHSSRLADH 180 ELKEMLLKKY SGCLSRLRSE FLKKRKKGKL PKDARTVLLE WWNTHYRWPY PTEEDKVRLA 240 AMTGLDPKQI NNWFINQRKR HWKPSEDMRF ALMEGVAGGS SGTTLYFDTG TIGP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 196 | 205 | LRSEFLKKRK |
| 2 | 202 | 206 | KKRKK |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Highly expressed in the early globular stage embryo before 2 days after pollination (DAP), but not in the endosperm. At 3 and 4 DAP, expression is restricted to the region around or just below the center of the ventral side of the embryo, where the shoot apex subsequently arises. During the transition to the shoot apex differentiation stage, expression is divided between the upper and basal regions of the shoot area, and the notch between the first leaf primordium and epiblast, respectively. When the first leaf primordia is evident, expression is localized to the notches between the shoot apical meristem (SAM) and the first leaf primordium and the putative second leaf primordium. Expressed uniformly in the inflorescence meristem, but after the transition from inflorescence to the floral phase, located specifically in the notches between the floral meristem and glume primordia. At later stages of flower development, uniformly expressed throughout the corpus of the meristem, and in the notches between glume primordia, but less well defined than in the previous stage. {ECO:0000269|PubMed:10488233}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
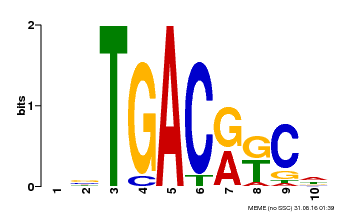
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.003G144200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY781901 | 0.0 | AY781901.1 Saccharum hybrid cultivar Pindar isolate c53a homeobox protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002455513.1 | 0.0 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 1e-166 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | C5XI60 | 0.0 | C5XI60_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g012480.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 | Representative plant | OGRP167 | 17 | 148 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.1 | 1e-84 | KNOTTED1-like homeobox gene 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.003G144200.1.p |
| Entrez Gene | 8060322 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



