 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.002G141300.1.p | ||||||||
| Common Name | Sb02g013010, SORBIDRAFT_02g013010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 367aa MW: 38984.9 Da PI: 9.5495 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 60.5 | 3.5e-19 | 21 | 66 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+g+W++eEde l ++v+++G ++W++I+r ++ gR++k+c++rw +
Sobic.002G141300.1.p 21 KGPWSPEEDEALQRLVARHGARNWSLISRSIP-GRSGKSCRLRWCNQ 66
79******************************.***********985 PP
| |||||||
| 2 | Myb_DNA-binding | 53.5 | 5.7e-17 | 75 | 117 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++T+eEd+ +++a++++G++ W+tIar + gRt++ +k++w++
Sobic.002G141300.1.p 75 PFTPEEDDTILRAHARFGNK-WATIARLLS-GRTDNAIKNHWNST 117
89******************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 26.597 | 16 | 71 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.12E-32 | 18 | 114 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.3E-17 | 20 | 69 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.4E-18 | 21 | 66 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-26 | 22 | 74 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.21E-16 | 23 | 65 | No hit | No description |
| SMART | SM00717 | 1.7E-14 | 72 | 120 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.338 | 73 | 122 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.9E-14 | 75 | 117 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-23 | 75 | 121 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.46E-11 | 75 | 118 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
MASSSPSCRG GGNKADVDRI KGPWSPEEDE ALQRLVARHG ARNWSLISRS IPGRSGKSCR 60 LRWCNQLSPQ VEHRPFTPEE DDTILRAHAR FGNKWATIAR LLSGRTDNAI KNHWNSTLKR 120 KYYAASAADG DAANGGAADA DDERPLKRTS SDGHPGLCFS PGSPSGSDLS DSSHHSLPSV 180 MPSAASAAAA AVTSQQQQQQ HVYRPVPRAG GVVVLPVAPP LAPRPPSPPP PPPPQQQPPP 240 ATSLSLSLSL PGLDQQPEPS PATAPPQPPV QIHQHQAAPP QMPPPPPQAQ PSPRLPFQLQ 300 PAPATNLTSP QPAAAAPFSS EFLLMMQEMI RIEVRNYMSG SGFDPRADGA SVHAVTKRMM 360 GMAKIE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-38 | 18 | 121 | 4 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.1608 | 0.0 | callus| embryo| leaf| shoot | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00476 | DAP | Transfer from AT4G37260 | Download |
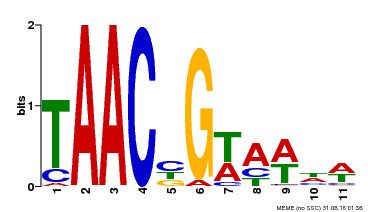
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.002G141300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HF679425 | 0.0 | HF679425.1 Saccharum hybrid cultivar Co 86032 mRNA for ScMYB19 protein. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002462029.1 | 0.0 | transcription factor MYB44 | ||||
| TrEMBL | C5X6Q8 | 0.0 | C5X6Q8_SORBI; Uncharacterized protein | ||||
| STRING | Sb02g013010.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP789 | 38 | 154 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37260.1 | 9e-64 | myb domain protein 73 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.002G141300.1.p |
| Entrez Gene | 8063857 |



