 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.002G050700.3.p | ||||||||
| Common Name | Sb02g004250, SORBIDRAFT_02g004250 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 764aa MW: 82165.9 Da PI: 6.4943 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 50.3 | 4.8e-16 | 449 | 487 | 2 | 40 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
+k+C+CkkskClk+YCeCfaag++Cse C+C++C Nk
Sobic.002G050700.3.p 449 ACKRCSCKKSKCLKLYCECFAAGVYCSEPCSCQGCLNKP 487
589**********************************96 PP
| |||||||
| 2 | TCR | 49.6 | 8e-16 | 534 | 573 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+gCnCkks+ClkkYCeC++ g+ Cs +C+Ce+CkN+
Sobic.002G050700.3.p 534 RHKRGCNCKKSSCLKKYCECYQGGVGCSTNCRCESCKNTF 573
589***********************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 3.8E-17 | 448 | 489 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 36.911 | 449 | 574 | IPR005172 | CRC domain |
| Pfam | PF03638 | 8.6E-12 | 451 | 486 | IPR005172 | CRC domain |
| SMART | SM01114 | 5.1E-16 | 534 | 575 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 1.6E-11 | 536 | 572 | IPR005172 | CRC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009934 | Biological Process | regulation of meristem structural organization | ||||
| GO:0048444 | Biological Process | floral organ morphogenesis | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 764 aa Download sequence Send to blast |
MNMQDSPVFS FLNNLSPIEP LKSAYNTNSL QGFQSINITS ISSIFTSPHD NVNKEPRLPK 60 SSLGDISENE VCADGADTNK PSKSSNAVRL FACTSTVTRE THSVTCSGVV DPPTGPCDLA 120 QPGQFDNGSP DHNTTPCHGV RSDLKLDKTR KLEAAQTVKN TLEKRKCLFS TDIQLLDGSS 180 EPVNDNDQVL GCEWSDLIST TSAELLAFDS TMDEHHRGVH LAAKNAESCG YLLSKLTGDG 240 DVSDRAHPSA SGQVYYQELV MGEDQTENAE IFQDGQETIS TEEIQDNICE ANGCLPLEYK 300 VEGQQQRGVR RRCLVFEASG FSNSVVKKGT VENLSVSTCK GKSPVQTQPR GLRGIGLHLN 360 ALALTPKGKM ACQDPTASAL LPSSASEKDV HGRLLSAGEN FTHSGGELLE FPVDDCSAGG 420 FPVNDHVSGQ SVSPQKKRRK TDNGDDGEAC KRCSCKKSKC LKLYCECFAA GVYCSEPCSC 480 QGCLNKPIHE EIVLSTRKQI EFRNPLAFAP KVIRMSDAGL ETGEDPNNTP ASARHKRGCN 540 CKKSSCLKKY CECYQGGVGC STNCRCESCK NTFGRRDAES ELTEEMKQES EQTENCSQEK 600 ENGQQKANVQ SEDHPLLDLV PITPPFDLSS SLLKLPNFSS AKPPRPSKPR SVNSRSSASK 660 ATATVQPCKS SKIAVSGIDE EMPDILKEPN SPSNCVKTTS PNGKRVSPPH NALSVSPNRK 720 GGRKLILKSI PSFPSLAGDT ISGSAMNNTD STFSVSPLTL GPS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 8e-15 | 451 | 571 | 12 | 120 | Protein lin-54 homolog |
| 5fd3_B | 8e-15 | 451 | 571 | 12 | 120 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 435 | 439 | KKRRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
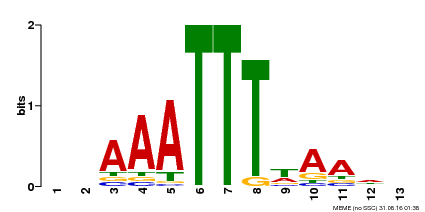
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.002G050700.3.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002459413.1 | 0.0 | protein tesmin/TSO1-like CXC 3 | ||||
| TrEMBL | A0A1W0W2A9 | 0.0 | A0A1W0W2A9_SORBI; Uncharacterized protein | ||||
| STRING | Sb02g004250.1 | 0.0 | (Sorghum bicolor) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14770.1 | 2e-77 | TESMIN/TSO1-like CXC 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.002G050700.3.p |
| Entrez Gene | 8081186 |



