 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.001G451000.1.p | ||||||||
| Common Name | Sb01g042400, SORBIDRAFT_01g042400 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 687aa MW: 73679.8 Da PI: 6.734 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 89.9 | 2.2e-28 | 201 | 254 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av++L G++kA+Pk+ilelm+v+gLt+e+v+SHLQk+Rl
Sobic.001G451000.1.p 201 KPRVVWSVELHQQFVNAVNHL-GIDKAVPKKILELMNVPGLTRENVASHLQKFRL 254
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 77.5 | 4.9e-26 | 17 | 125 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedal 87
vl+vdD+p+ + +l+++l + +y +v+++ +++ al++l+e++ +D+i+ D+ mp+mdG++ll+ e++lp+i+++a + + +
Sobic.001G451000.1.p 17 VLVVDDDPTCLLVLKRMLLDCRY-DVTTCPQATRALTMLRENRrgFDVIISDVHMPDMDGFRLLELVGL-EMDLPVIMMSADSRTDIVM 103
89*********************.***************99999********************87754.558**************** PP
HHHHTTESEEEESS--HHHHHH CS
Response_reg 88 ealkaGakdflsKpfdpeelvk 109
+ +k Ga d+l Kp+ +eel +
Sobic.001G451000.1.p 104 KGIKHGACDYLIKPVRMEELKN 125
*******************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.7E-180 | 1 | 669 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 2.9E-42 | 14 | 150 | No hit | No description |
| SuperFamily | SSF52172 | 2.28E-36 | 14 | 139 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 4.1E-30 | 15 | 127 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.374 | 16 | 131 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 5.6E-23 | 17 | 125 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.00E-27 | 18 | 130 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-29 | 198 | 259 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.674 | 198 | 257 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.14E-20 | 198 | 258 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.0E-25 | 201 | 254 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.8E-7 | 203 | 252 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 687 aa Download sequence Send to blast |
MAAAEARGGE FPVGMKVLVV DDDPTCLLVL KRMLLDCRYD VTTCPQATRA LTMLRENRRG 60 FDVIISDVHM PDMDGFRLLE LVGLEMDLPV IMMSADSRTD IVMKGIKHGA CDYLIKPVRM 120 EELKNIWQHV VRKKFNENKD HEHSGSLDDT DRNRPTNNDN EYASSANDGG DGSWKSQKKK 180 REKEDDETDL ENGDPSSTSK KPRVVWSVEL HQQFVNAVNH LGIDKAVPKK ILELMNVPGL 240 TRENVASHLQ KFRLYLKRIA QHHAGIPHPF VAPASSAKVA PLGGLEFQAL AASGQIPPQA 300 LAALQDELLG RPTSSLALPG RDQSSLRLAA IKGNKSHGER EIAFGQPIYK CQNNAYGAFP 360 QSSPSVGGLP SFAAWPNNKL GMTDSPSTLG NVGSSQNNNM LLHELQQQPD TLLSGTLHNI 420 DVKPSGVVMP SQSLNAFPAS EGISPNQNPL ILPSQSSSFL ASIPPSMKHE PLLGSPSPST 480 SLLGGLDMVN QASTSQPLIS SHGANLPGLI NRSTNAMPSP GINNFQSGNI PYLVNQNAMG 540 VSSRPPGVLK TESTDSLSRS YGYIGGSTSV DSVLLSSQSK NAQYGLLQSQ NDVSGSWSPS 600 QDFDSFGNSL GQGHPGSTSS NFQSSALGKL PDQGRGRNHG FVGKGTCIPS RFAVDEVESP 660 TNNLSHSIGN SGDIVNPDIF GFSGQM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-24 | 197 | 260 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.12399 | 0.0 | callus| leaf | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
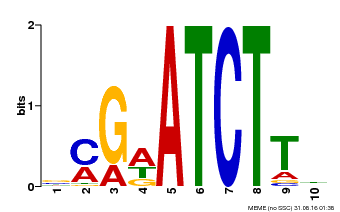
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.001G451000.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB060130 | 0.0 | AB060130.1 Zea mays ZmRR8 mRNA for response regulator 8, complete cds. | |||
| GenBank | EU975545 | 0.0 | EU975545.1 Zea mays clone 492147 two-component response regulator ARR1 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002468245.1 | 0.0 | two-component response regulator ORR21 | ||||
| Refseq | XP_021306470.1 | 0.0 | two-component response regulator ORR21 | ||||
| Swissprot | A2XE31 | 0.0 | ORR21_ORYSI; Two-component response regulator ORR21 | ||||
| TrEMBL | C5WSM4 | 0.0 | C5WSM4_SORBI; Two-component response regulator | ||||
| STRING | Sb01g042400.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5796 | 37 | 54 | Representative plant | OGRP292 | 17 | 120 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-114 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.001G451000.1.p |
| Entrez Gene | 8056866 |



