 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_04734.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 377aa MW: 41226.3 Da PI: 10.1683 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.8 | 1.1e-12 | 68 | 116 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++kGV + +grW A+I++ ++r++lg+f +eeAa+ ++ a+++++g
Rsa1.0_04734.1_g00001.1 68 SKFKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNEEEEAAASYDIAARRFRG 116
79****9888.8*********3.....5*************************98 PP
| |||||||
| 2 | B3 | 102.9 | 1.7e-32 | 201 | 312 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT...............---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHH CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh...............ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvka 71
f+k+ tpsdv+k++rlv+pk++ae+h + ++++++ + led++g++W+++++y+++s++yvltkGW++Fvk+
Rsa1.0_04734.1_g00001.1 201 FEKAVTPSDVGKLNRLVIPKQHAEKHfplpamttatavtamSPSPTKGVLINLEDRTGKVWRFRYSYWNSSQSYVLTKGWSRFVKE 286
89******************************999999998667789*************************************** PP
HT--TT-EEEEEE-SSSEE..EEEEE CS
B3 72 ngLkegDfvvFkldgrsefelvvkvf 97
++L++gD+v F+++ +++l++ ++
Rsa1.0_04734.1_g00001.1 287 KNLRAGDVVCFERSTGPDRQLYIDWR 312
************88778888888886 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.26E-24 | 68 | 123 | No hit | No description |
| Pfam | PF00847 | 8.6E-7 | 68 | 116 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 3.4E-16 | 68 | 124 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 19.454 | 69 | 124 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.5E-19 | 69 | 124 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.9E-27 | 69 | 130 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.1E-40 | 195 | 314 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.45E-31 | 199 | 303 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.16E-29 | 200 | 300 | No hit | No description |
| SMART | SM01019 | 1.5E-25 | 201 | 315 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.944 | 201 | 315 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.3E-29 | 201 | 312 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 377 aa Download sequence Send to blast |
MDNSCIDDST SNNTSGSLSI STPSAKKKLS PPPPQATMRL YRMGSGGSSV VLDSENGVET 60 ESRKLPSSKF KGVVPQPNGR WGAQIYEKHQ RVWLGTFNEE EEAAASYDIA ARRFRGRDAV 120 TNFKSPPATT ALDGAADAES AFLEAHSKAE IVDMLRKHTY ADELQQSKRK FLDGNGNGKS 180 GTAMSNSRGN DGVSRAREVL FEKAVTPSDV GKLNRLVIPK QHAEKHFPLP AMTTATAVTA 240 MSPSPTKGVL INLEDRTGKV WRFRYSYWNS SQSYVLTKGW SRFVKEKNLR AGDVVCFERS 300 TGPDRQLYID WRVQSGTEGI TPVRSVVRLF GVNISNAATN AKSNDVAAEY GGGKKRSREV 360 DLFASGCSKK QTMINAL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 2e-57 | 198 | 318 | 11 | 121 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
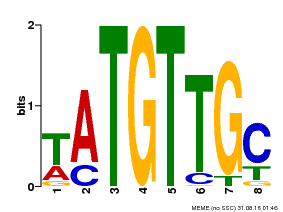
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_04734.1_g00001.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018458192.1 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| Swissprot | Q9C6M5 | 0.0 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A0A0D3CD61 | 0.0 | A0A0D3CD61_BRAOL; Uncharacterized protein | ||||
| STRING | Bo5g043120.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 0.0 | RAV family protein | ||||



