 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00782.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 995aa MW: 111093 Da PI: 6.0784 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 189 | 4.6e-59 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYa 86
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYa
Rsa1.0_00782.1_g00001.1 19 LSEaQHRWLRPAEICEILRNYQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYA 104
45559********************************************************************************* PP
CG-1 87 hseenptfqrrcywlLeeelekivlvhylevk 118
h+e+n++fqrrcyw+Le+el +iv+vhylevk
Rsa1.0_00782.1_g00001.1 105 HGEDNENFQRRCYWMLEQELMHIVFVHYLEVK 136
*****************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 86.826 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 8.0E-87 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.5E-52 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 5.53E-13 | 405 | 489 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 5.44E-10 | 585 | 687 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 10.365 | 587 | 636 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.6E-10 | 587 | 688 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.80E-7 | 593 | 685 | No hit | No description |
| PROSITE profile | PS50088 | 9.351 | 604 | 636 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 803 | 825 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.12E-7 | 804 | 853 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0019 | 805 | 824 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.858 | 805 | 833 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 826 | 848 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 827 | 851 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.4E-4 | 829 | 848 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 995 aa Download sequence Send to blast |
MADRGSFGFA PQLDIQQLLS EAQHRWLRPA EICEILRNYQ KFHIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEDNE NFQRRCYWML 120 EQELMHIVFV HYLEVKGNRI SSSGTKENNS NSLSGSTSVN IESTANTSST LSPLCEDADS 180 GNRDGWASQV HGNRVKESDS QRLVGVPALD ASFENPLARF QNLPPYNNAL LTQTNPSNTG 240 LMSVEGHLRS PLHNQVNWQI PVQDSLPLQK WPPMDSHSGM AHGTDLTLHE TFSSLIGSQN 300 QQPSFQAPFT SVETAYIPKF GPEDLLYEAS ANQTLPLRKS LLKKEDSLKK VDSFSRWVSN 360 ELAEMEDLQM QSSSGGGIAW TSVETAAAAS SLSPSLSKDQ RFTMIDFWPK WTQTDSEVEV 420 MVIGTFLLSP QEVTSYSWAC MFGEVEVPAE ILVDGVLCCH APPHEVGQVP FYITCSDRFS 480 CSEVREFDFL PGSARKLNAA DLYGAFTNEA SLHLRFENLL SRVSSSAQEH NIFEDVGEKR 540 RKISRIMLLK DEKESLLAST AEKGLTEVEA KERLIREEFE DKLYLWLIHK VTEEGKGPNI 600 LDEEGQGVLH LAAALGYDWA IKPILAAGVS INFRDANGWE DTVALLVSLG ADSGALTDPS 660 PELPLGKTAS DLAYGNGHRG ISGFLAESSL TSYLEKLTVD GKEDSSTDSS RAKAVQTVAE 720 RTATPMSYGD VPETLSMKDS LTAVLNATQA ADRLHQVFRM QSFQRKQMSE IGEKNEFGVS 780 DELAVCFAAG KSKKAGHSSG GAAVHAAAVQ IQKKYRGWKK RKEFLLIRQR IVKIQAHVRG 840 HQVRKQYRAI IWSVGLLEKI ILRWRRKGSG LRGFKRDAVT KAPEPVCAAP TQEDDYDFLK 900 EGRKQTEERL QKALTRVKSM AQYPEARAQY RRLLTVVEGI RENEASSSSA MNNNNNNNTN 960 TNSNTEEAAN YNEEDDLIDI DSLLDDDTFM SLAFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
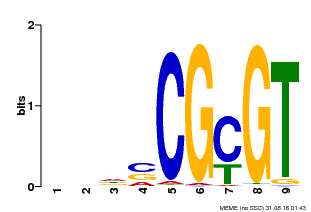
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00782.1_g00001.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018456913.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A397XS62 | 0.0 | A0A397XS62_BRACM; Uncharacterized protein | ||||
| STRING | Bo9g018010.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



