 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00589.1_g00010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 342aa MW: 39154.4 Da PI: 7.4167 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.3 | 6.2e-32 | 60 | 113 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+e+AtPk ++++m++kgL+++hvkSHLQ+YR+
Rsa1.0_00589.1_g00010.1 60 PRLRWTPDLHLRFVRAVERLGGQERATPKLVRQMMNIKGLSIAHVKSHLQMYRS 113
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.273 | 56 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.6E-29 | 56 | 114 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.97E-14 | 58 | 114 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-22 | 60 | 115 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.2E-9 | 61 | 112 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MIPSIEGSGN TDREEEEEEE EEEEEEDGEG EESKISSNTT VEAEVNKKTK VRPYVRSKVP 60 RLRWTPDLHL RFVRAVERLG GQERATPKLV RQMMNIKGLS IAHVKSHLQM YRSKKMDDQG 120 QAIADHRHFM ETSTDRNIYK LSQLPMFRGY NMNYNHDSPF RYGSKFSNAS LWSSSSHETN 180 RSLIDRPGSI CGSSVSNNIH GSEYLTNNRS FQNNFSSKIS NHLPKLRYNH QERTNLATFN 240 SIQGHSRAFQ KFETGNEERT NHAYCNKTAG KRNASTRIDL DLSLKLRVPE ETVLEETETA 300 ATTTDQTLSL SLCSWKKSRV IKTDEEDRTV KNGQASTLDL TL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 8e-17 | 61 | 115 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-17 | 61 | 115 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-17 | 61 | 115 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-17 | 61 | 115 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
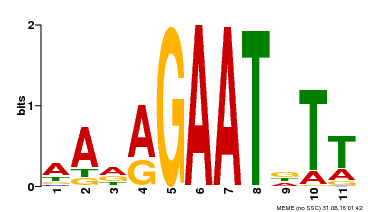
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00589.1_g00010.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018437061.1 | 0.0 | PREDICTED: myb family transcription factor PHL8 | ||||
| TrEMBL | A0A0D3BQY8 | 0.0 | A0A0D3BQY8_BRAOL; Uncharacterized protein | ||||
| STRING | Bo4g029640.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM363 | 28 | 184 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 1e-173 | G2-like family protein | ||||



