 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00547.1_g00003.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 970aa MW: 107232 Da PI: 5.5339 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.8 | 5e-29 | 597 | 647 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
Rsa1.0_00547.1_g00003.1 597 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 647
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF55781 | 4.76E-5 | 214 | 281 | IPR029016 | GAF domain-like |
| PROSITE profile | PS51519 | 17.393 | 586 | 667 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.4E-26 | 599 | 647 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.57E-23 | 867 | 958 | No hit | No description |
| Pfam | PF00564 | 3.4E-20 | 874 | 955 | IPR000270 | PB1 domain |
| SMART | SM00666 | 5.2E-29 | 874 | 956 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 6.6E-27 | 874 | 956 | No hit | No description |
| PROSITE profile | PS51745 | 28.012 | 874 | 956 | IPR000270 | PB1 domain |
| CDD | cd06407 | 5.78E-35 | 875 | 955 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 970 aa Download sequence Send to blast |
MCEPDDNTTA RNGVTPPPSR SRELLMDVDD LDLDGSWPLD QIPYLSSSNR MISPIFVSSS 60 SEQPCSPLWA FSDAVGGTVN HGGDDEKISS ASGVPSFRLS DYPLFLPYSS SPSVAENTTE 120 KHNGFQFPSP LMSLVPPENT DNYCVIKERM TQALRYFKES TEQHVLAQVW APVRKNGRDL 180 LTTLGQPFVL NPNGNGLNQY RMISLTYMFS VDSESDFELG LPGRVFRQKL PEWTPNVQYY 240 SSKEFSRLDH ALHYNVRGTL ALPVFNPSGE SCIGVVELIM TSEKIHYAPE VDRVCKALEF 300 NKIWSFWQAV NLKSSEILDH QTTQICNESR QNALAEILEV LTVVCETHNL PLAQTWVPCQ 360 HGSVLANGGG LKKNCTSFDG SCMGQVCMST TDMACYVVDA HVWGFRDACL EHHLQKGQGV 420 AGRAFLNGGS CFCRDITKFC KTQYPLVHYA LMFKLTTCFA ISLQSSYTGD DSYILEFFLP 480 SSITDNQEQD SLLGSLLVTM KEHFQSLRVA SGVDFGEDDD KLSFEIIEAL PDKKIHSQIE 540 SIRVPFSGFK SNATETLLIP QPAVPSSDHV NGKTNVAPVN GVVKEKKKTE KKRGKTEKTI 600 SLDVLQQYFT GSLKDAAKSL GVCPTTMKRI CRQHGISRWP SRKIKKVNRS ITKLKRVIES 660 VQCTDGGLDL TSMAVSSIPW THGQTAAQQL NSPSGSKPPE LPTTNNSPNH WSNEHSPPEP 720 NGSPELPSNG HKRSRTADES AGTPTSHGSC DGNGNQLDET KVPSQDPLFT VGGGPPGLFF 780 PPYSRDHDVS AASFAMPNRL LGTIDHFRGM PIEDAGSSKD LRNLCSTAAF EDKFPDSNWM 840 NNDNNSNNNM YVPPKEEEVA NITREPPSES EMRTVTIKAS YKEDIIRFRI PSGSGITELK 900 DEVAKRLRID AGTFDIKYLD DDNEWVLIAC DADLQECLEI PRSSRTNIVR LLVHDVTANL 960 GSSCESTGEL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
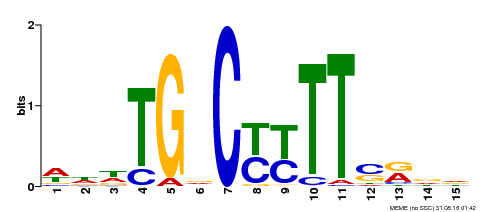
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00547.1_g00003.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228452 | 0.0 | AK228452.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-03-P13. | |||
| GenBank | BT005776 | 0.0 | BT005776.1 Arabidopsis thaliana clone RAFL15-03-P13 (R20174) unknown protein (At4g24020) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018469935.1 | 0.0 | PREDICTED: protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | M4DB94 | 0.0 | M4DB94_BRARP; Uncharacterized protein | ||||
| STRING | Bra013754.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



