 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_00064.1_g00036.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 925aa MW: 102487 Da PI: 6.0508 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 135.3 | 1.8e-42 | 148 | 225 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+Ceadls++k+yhrrhkvCe+hska+++lv g++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Rsa1.0_00064.1_g00036.1 148 VCQVENCEADLSKVKDYHRRHKVCEMHSKATSALVGGIMQRFCQQCSRFHVLEEFDEGKRSCRRRLAGHNKRRRKTNP 225
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.6E-34 | 142 | 210 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.582 | 146 | 223 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.62E-39 | 148 | 227 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.3E-31 | 149 | 222 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.48E-8 | 716 | 815 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.1E-8 | 717 | 821 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.91E-9 | 718 | 815 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 925 aa Download sequence Send to blast |
MEARIEGEGQ HFYGYRAMDV RSVGKRSVEW DLNDWKWDGD LFVATPLNPG ASRQFFPLGN 60 SSNSSSTCSE EGNNNNSREV EKKRRAPASA ATATGDDDDN TNNNGGGGLT LKIGENGYDL 120 NGEREGNSAA AKKTKFGGGG GGVANRSVCQ VENCEADLSK VKDYHRRHKV CEMHSKATSA 180 LVGGIMQRFC QQCSRFHVLE EFDEGKRSCR RRLAGHNKRR RKTNPEPGAN GNPLSDDHQS 240 SNLLLITLLK ILSNMHSNGS DHPGDQDLMP HLLKSLVSHA GEQLGKNLVE LLLQGGGSQA 300 SGLLAVEQAP QEDLKQAPQL PRQELYANGT ATGNRSEKQA RVNDFDLNDI YIDSDDGTDI 360 ERSSPPPTNP ATSSPDYPSW IHQTSPPQTT RNSDSASDQS PSSSSEDAQM RTGRIVFKLF 420 GKEPNDFPVV LRGQILDWLS HTPTDIESYI RPGCIVLTIY LRQAETAWDE LSDDMGFSLS 480 KLLDLSDDPL WTTGWIYVRV QNQYAFVFDG QVAVDTSLPL RSRDYSHITS VRPLAVAATG 540 KAQFTVKGIN LRRPGTRLLC AVEGKYLIQE TTKEDDDLKE SNEIVECVSF SCDLPITSGR 600 GFMEIEDQGL SSSFFPFIVV EEDDVCSEIR VLETTLEFTG TDSAKQAMEF IHELGWLLQR 660 SKLGVFPLTR FKWLIEFSMD REWCAVIRKL LNMFFEGAVG DYSSSSPSDA ALSELCLLHR 720 AVRKNSKPMV EMLLRYSPKQ QRIYSLFRPD AAGPAGLTPL HIAAGKDGSE DVLDALTEDP 780 AMVGVGAWRT SRDSTGFTPE DYARLRGHFS YIHLIQRKIN KKSATEDHVV VNIPASISDR 840 EQKETKPGSS ALEITHGNHK LQCKLCDHKL VYGTGRRSVA YRPAMLSMVA IAAVCVCVAL 900 LFKSCPEVLY VFQPFRWELL DYGTR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-32 | 139 | 222 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
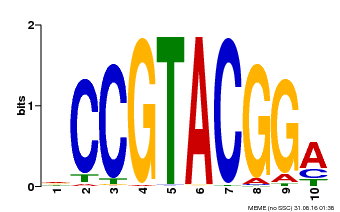
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_00064.1_g00036.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC815948 | 0.0 | KC815948.1 Arabis alpina SQUAMOSA promoter binding protein-like protein 1 (SPL1) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018439364.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Refseq | XP_018439365.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A1J3IQZ6 | 0.0 | A0A1J3IQZ6_NOCCA; Squamosa promoter-binding-like protein 1 | ||||
| STRING | XP_006397898.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3030 | 27 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



