 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC951_p3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 419aa MW: 47349.1 Da PI: 10.4121 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 30.2 | 9.5e-10 | 131 | 171 | 5 | 45 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka++++Le+ +L++ ++L
RrC951_p3 131 KTVRRLAQNREAARKSRLRKKAYVQQLENSRLKLTQLEQEL 171
7789************************9877777766555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.5E-7 | 127 | 190 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.438 | 129 | 173 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 9.0E-7 | 131 | 171 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 5.3E-8 | 131 | 176 | No hit | No description |
| SuperFamily | SSF57959 | 5.27E-7 | 131 | 174 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 134 | 149 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 4.2E-33 | 218 | 292 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 419 aa Download sequence Send to blast |
RTKTCRRLDI RILSLSLSLF SLLLGFKSDY KAQPFLQWTF PLQHTKRCSF FKFQSVHTRE 60 VSRVLYCKRG RHKWILSLAI KCVSTEASMA DMSPITDVST DGDRDLGSEG ALVNTAASDS 120 SERSKNKLDQ KTVRRLAQNR EAARKSRLRK KAYVQQLENS RLKLTQLEQE LQRARQQGVF 180 ISSTGDQAHS TGGNGWLNRA LAFDAEYSRW LEEKNKRMNE LRSALNAHAG DGELRIIVDG 240 VMAHYEELFR IKSNAAKNDV FHLLYGMWKT PAERCFLWLG GFRSSEILKL LANQLEPMTE 300 RQVMGINSLQ ETSQQAEDAL SQGMESLQQS LADTLSSGTL GSSSSGNVAS YMGQMAMAMG 360 KLGTLEGFIR QADNLRLQTL QQMIRVLTTR QSARALLAIH DYSSRLRALS SLWLARPRE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. May be involved in the induction of the systemic acquired resistance (SAR) via its interaction with NPR1. Could also bind to the Hex-motif (5'-TGACGTGG-3') another cis-acting element found in plant histone promoters (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
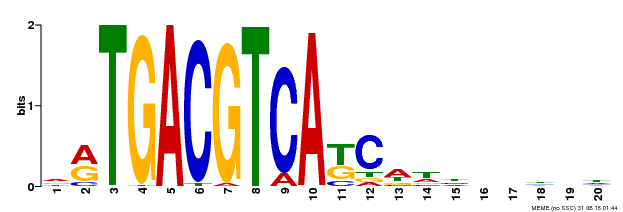
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353125 | 0.0 | AK353125.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-10-O14. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018445153.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Refseq | XP_018445154.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Refseq | XP_018445155.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Refseq | XP_018445327.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Refseq | XP_018445328.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Refseq | XP_018445329.1 | 0.0 | PREDICTED: transcription factor TGA2-like | ||||
| Swissprot | Q39140 | 0.0 | TGA6_ARATH; Transcription factor TGA6 | ||||
| TrEMBL | A0A3P6DNB4 | 0.0 | A0A3P6DNB4_BRACM; Uncharacterized protein | ||||
| STRING | Bo9g175710.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM667 | 28 | 133 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G12250.2 | 0.0 | TGACG motif-binding factor 6 | ||||



