 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC8582_p1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 407aa MW: 42833.9 Da PI: 9.3388 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 121.3 | 3.5e-38 | 91 | 150 | 3 | 62 |
zf-Dof 3 ekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkkss 62
e+alkcprCdstntkfCy+nny+l+qPr+fCkaCrryWt+GGalr vPvGgg+r+nk+++
RrC8582_p1 91 ETALKCPRCDSTNTKFCYFNNYNLTQPRHFCKACRRYWTRGGALRSVPVGGGCRRNKRTK 150
6789*****************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 7.0E-36 | 74 | 143 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 5.6E-32 | 93 | 148 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 29.553 | 94 | 148 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 96 | 132 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 407 aa Download sequence Send to blast |
MVFSSFPNYS DSSNWQQQHQ PITTTVGFTG NNISQQFLPH HPLPPQPQQT PPPLHHNGNG 60 GGGGGPGGPG GLIRPGSMAE RARLANIPLP ETALKCPRCD STNTKFCYFN NYNLTQPRHF 120 CKACRRYWTR GGALRSVPVG GGCRRNKRTK NSGGGGGSSS SGNSKSQDST TSNDQYHHRA 180 LSNNQMGPPS SSTSLSSLLS SYNAGLIPGH DHNSNNNNIL GLGSSLSPLK LMHPLDFTDN 240 FTLQYGAVSA PSYHTGGGSS GGNGGGGASA ILTGLDQWRF PATHHQLPLL GGLDSSSSSG 300 LYPFDQNPGY GLVTGSGQYR PKNIFHNLVS SSSSSSASSA MVTATASQLA SVKMEDSNNQ 360 LNMSRQLFGN EQELWNIHGT AVSTAAATTS SWSDVSNNFS SSSTSNI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Binds to 5'-TAAAGT-3' motif in REV promoter to triggers its transcription, thus regulating adaxial-abaxial polarity and influencing leaf axial patterning in an auxin transport- and response-dependent manner (e.g. IAA6 and IAA19 genes expression) (PubMed:20807212). Probably involved in early processes for vascular development (PubMed:17583520). The PEAR proteins (e.g. DOF2.4, DOF5.1, DOF3.2, DOF1.1, DOF5.6 and DOF5.3) activate gene expression that promotes radial growth of protophloem sieve elements (PubMed:30626969). {ECO:0000250|UniProtKB:Q9M2U1, ECO:0000269|PubMed:17583520, ECO:0000269|PubMed:20807212, ECO:0000269|PubMed:30626969}. | |||||
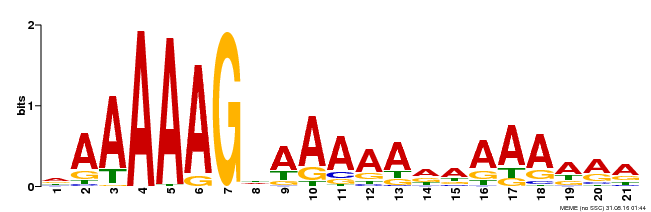
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00483 | DAP | Transfer from AT5G02460 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cytokinin in procambium. {ECO:0000269|PubMed:30626969}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC189646 | 0.0 | AC189646.2 Brassica rapa subsp. pekinensis cultivar Inbred line 'Chiifu' clone KBrS009G14, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018453553.1 | 0.0 | PREDICTED: dof zinc finger protein DOF5.1-like | ||||
| Swissprot | Q9LZ56 | 1e-166 | DOF51_ARATH; Dof zinc finger protein DOF5.1 | ||||
| TrEMBL | A0A3P5ZX96 | 0.0 | A0A3P5ZX96_BRACM; Uncharacterized protein | ||||
| STRING | Bra028872.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM11519 | 17 | 32 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G02460.1 | 1e-126 | Dof family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



