 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC700_p7 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 261aa MW: 27524.5 Da PI: 7.8383 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 124.8 | 9.5e-39 | 31 | 116 | 3 | 87 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec.eaesssssasnsss 87
+k+++++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti+WLlqqa+p+i+++tg+++ +as+ +a + +s++++++
RrC700_p7 31 PKRSSNKDRHTKVEGRGRRIRMPALCAARIFQLTRELGHKSDGETIQWLLQQAEPSIIAATGSGTVPASALaSACAAVSNQHHQGG 116
6999***********************************************************99999777666666666665554 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 3.6E-31 | 35 | 112 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 28.016 | 36 | 90 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 261 aa Download sequence Send to blast |
MDPKNPNQHH HHQIVVASDK EPNNSKKQLG PKRSSNKDRH TKVEGRGRRI RMPALCAARI 60 FQLTRELGHK SDGETIQWLL QQAEPSIIAA TGSGTVPASA LASACAAVSN QHHQGGSLTA 120 GLMISHHDLD GGGGRPSWGE GGGEASRLWP NGAGGVMSEG AGYRIGFPGF DFPGGAMSFA 180 SILGGGGNSI QIPGLELGLS QEGNVGVLNP QCFTHIYQQM AQAQALAQGR TVHHSLHHNP 240 SLEDHQQESG DQKVDSQGSG R |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the site II motif (3'-TGGGCC/T-5') in the promoter of PCNA-2 and to 3'-GCCCG/A-5' elements in the promoters of cyclin CYCB1-1 and ribosomal protein genes. {ECO:0000269|PubMed:12631321, ECO:0000269|PubMed:16123132}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00005 | PBM | Transfer from 484486 | Download |
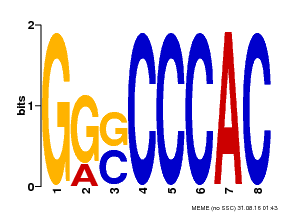
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018456599.1 | 0.0 | PREDICTED: transcription factor TCP20 | ||||
| Swissprot | Q9LSD5 | 1e-102 | TCP20_ARATH; Transcription factor TCP20 | ||||
| TrEMBL | A0A3P5YBZ8 | 1e-115 | A0A3P5YBZ8_BRACM; Uncharacterized protein | ||||
| STRING | XP_006395493.1 | 1e-112 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4050 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27010.1 | 4e-95 | TCP family protein | ||||



