 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC332_p4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 987aa MW: 111444 Da PI: 5.9021 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.7 | 7.4e-56 | 5 | 128 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvgg......vevlycyYahseenptf 94
l+e ++rwl++ ei++iL n++k+++++e++trp sgsl+L++rk++ryfrkDG++w+kkkdgkt++E+hekLKv + ++vl+cyYah+e n++f
RrC332_p4 5 LSEaQHRWLRPAEICEILRNYHKFHIATESPTRPASGSLFLFDRKVLRYFRKDGHNWRKKKDGKTIKEAHEKLKVRTlfevgsIDVLHCYYAHGEGNENF 104
45559***********************************************************************9999999***************** PP
CG-1 95 qrrcywlLeeelekivlvhylevk 118
qrrcyw+Le+el +iv+vhylevk
RrC332_p4 105 QRRCYWMLEQELMHIVFVHYLEVK 128
*********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.253 | 1 | 133 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.5E-80 | 4 | 128 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.4E-47 | 7 | 127 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 6.12E-14 | 396 | 482 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 18.669 | 579 | 661 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 9.5E-7 | 579 | 656 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.04E-17 | 580 | 690 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.67E-12 | 585 | 688 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-16 | 587 | 692 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0029 | 629 | 658 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.22 | 629 | 661 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 1.2 | 805 | 827 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 7.43E-8 | 805 | 855 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 8.004 | 806 | 835 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 8.3E-4 | 807 | 826 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0029 | 828 | 850 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.322 | 829 | 853 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 831 | 850 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 987 aa Download sequence Send to blast |
MEQLLSEAQH RWLRPAEICE ILRNYHKFHI ATESPTRPAS GSLFLFDRKV LRYFRKDGHN 60 WRKKKDGKTI KEAHEKLKVR TLFEVGSIDV LHCYYAHGEG NENFQRRCYW MLEQELMHIV 120 FVHYLEVKGS RTSIGMKENN SNSLSGTVSV NIDSAASPTS RLSSYCEDAD SVFGAGDSHQ 180 SSSVLQASPE PQTGNHNGWT SAPGMRSDSQ RLFDVQAWDA VGNLVTRYDQ PCNNLLVEER 240 TDKGGMLAAE HLRSPLQTQL NWQIPAQDDL PLPKWHLAPH SGMADDTDLA LFEQSAQDNF 300 ESFSSLLDIE HLQSDGIPPS NMETEFIPVK KSLLRHEDSL KKVDSFSRWA SKELGEMEDL 360 QMQSSRGDIA WTTVDCETAA AGLAFSPSLS EDQRFTIVDY WPKCAQTDAE VEVLVIGTFL 420 LSPQEVTICS WSCMFGEVEV PAEILVDGVL CCHAPPHTAG QVPFYVTCSN RFACSELREF 480 EFLSGSTKKI DAADIYGYSS KEASLQLRFE KLLARRDFVQ EHQIFEDVVE KRRKISSIML 540 LNEEKEYIFP GMYERDSTKQ EPKERVFREQ FEDELYIWLI HKVTEGGKGP NMLDEGGQGV 600 LHFVAALGYD WAIKPILAAG VNINFRDSNG WSALHWAAFS GREETVAVLV SLGADAGALT 660 DPSPELPLGK TAADLAYGKE HRGISGFLAE SSLTSYLEKL TMVSKENSPA NSSGPKAVQT 720 VSERTAAPMS YGDVPEKLSM KDSLTAVRNA TQAANRLHQV FRMQSFQRKQ LSGFDDDDDE 780 LGISDELAVS FAASKTKNPG HSEVFAHSAA THIQKKYRGW KKRKEYLLIR QRVVKIQAHV 840 RGHQVRKQYK PIVWSVGLLE KIILRWRRKG TGLRGFKRNA VPKTVEPEPL SPMMIPKEDD 900 YDFLDKGRKQ TEERLQKALT RVKSMVQYPE ARDQYRRLLT VVEGFRENEA SSSLSINNRE 960 ELVNCEEDDL IDIESLLNDD ILMSTSP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds calmodulin in a calcium-dependent manner in vitro (PubMed:11925432). Binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance (PubMed:19270186). Involved in freezing tolerance in association with CAMTA2 and CAMTA3. Contributes together with CAMTA2 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved in drought stress responses by regulating several drought-responsive genes (PubMed:23547968). Involved in auxin signaling and responses to abiotic stresses (PubMed:20383645). Activates the expression of the V-PPase proton pump AVP1 in pollen (PubMed:14581622). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:11925432, ECO:0000269|PubMed:14581622, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:23547968, ECO:0000269|PubMed:23581962, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
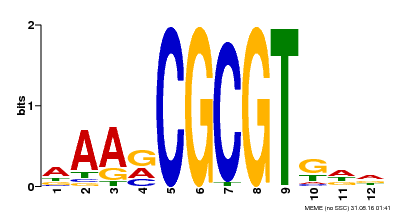
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by UVB, wounding, ethylene and methyl jasmonate (PubMed:12218065). Induced by salt stress and heat shock (PubMed:12218065, PubMed:20383645). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:20383645, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF491304 | 0.0 | AF491304.1 Brassica napus calmodulin-binding transcription activator (CAMTA) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018446094.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 1 isoform X2 | ||||
| Refseq | XP_018446095.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 1 isoform X2 | ||||
| Swissprot | Q9FY74 | 0.0 | CMTA1_ARATH; Calmodulin-binding transcription activator 1 | ||||
| TrEMBL | M4CYT5 | 0.0 | M4CYT5_BRARP; Uncharacterized protein | ||||
| STRING | Bra009382.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 0.0 | ethylene induced calmodulin binding protein | ||||



