 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC272_p5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 317aa MW: 35157.6 Da PI: 9.8825 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 45 | 2.6e-14 | 64 | 108 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
RrC272_p5 64 RENWTEQEHDKFLEALHLFDRD-WKKIEAFVG-SKTVVQIRSHAQKY 108
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.97E-17 | 58 | 114 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.6 | 59 | 113 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.6E-7 | 61 | 114 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.2E-18 | 62 | 111 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.3E-11 | 63 | 111 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-12 | 64 | 108 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.74E-9 | 66 | 109 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MVSLNARPAP FFDSVSMSLP GLDSRASDSD DGFGLTPIAR RDAASFREDT TAKIRKPYTI 60 KKSRENWTEQ EHDKFLEALH LFDRDWKKIE AFVGSKTVVQ IRSHAQKYFL KVQKSGANEH 120 LPPPRPKRKA SHPYPHKAPK NFALPSHAVG SLPASSAPLV QPAYLYSSSS DSRSLVGNPP 180 PASSSSSSQN HALTKPVIQE EPGVSATALP KNRCCSTGNK EVTQRLPAVR ESNDDDDDED 240 SCGKPRRVMP NFAEVYRFIG SVFDPNTSGH LQRLKQMDPI NIETVLLLMR NLSINLTSPE 300 FSEQRKLISS YSSKALK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
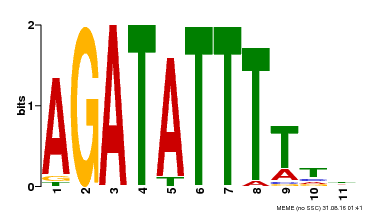
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00423 | DAP | Transfer from AT4G01280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ937212 | 3e-71 | AJ937212.1 Arabidopsis thaliana mRNA for myb transcription factor LHY-CCA1-like4 (lcl4 gene). | |||
| GenBank | AY519513 | 3e-71 | AY519513.1 Arabidopsis thaliana MYB transcription factor (At4g01280) mRNA, complete cds. | |||
| GenBank | BT024711 | 3e-71 | BT024711.1 Arabidopsis thaliana At4g01280 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018456205.1 | 0.0 | PREDICTED: protein REVEILLE 5 | ||||
| Swissprot | C0SVG5 | 1e-149 | RVE5_ARATH; Protein REVEILLE 5 | ||||
| TrEMBL | A0A397XRG4 | 1e-174 | A0A397XRG4_BRACM; Uncharacterized protein | ||||
| STRING | Bo9g003580.1 | 1e-170 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1290 | 28 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01280.1 | 1e-144 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



