 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC1548_p3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1007aa MW: 112391 Da PI: 6.0791 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 189 | 4.7e-59 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+e+n++fqrrcy
RrC1548_p3 19 LSEaQHRWLRPAEICEILRNYQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGEDNENFQRRCY 117
45559********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
w+Le+el +iv+vhylevk
RrC1548_p3 118 WMLEQELMHIVFVHYLEVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 86.826 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 8.0E-87 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.6E-52 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 5.6E-13 | 420 | 504 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 10.471 | 602 | 651 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.27E-9 | 602 | 702 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.6E-9 | 603 | 703 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 6.70E-7 | 608 | 700 | No hit | No description |
| PROSITE profile | PS50088 | 9.351 | 619 | 651 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 818 | 840 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.01E-7 | 819 | 868 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.858 | 820 | 848 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0019 | 820 | 839 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 841 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 842 | 866 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.4E-4 | 844 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1007 aa Download sequence Send to blast |
MADRGSFGFA PQLDIQQLLS EAQHRWLRPA EICEILRNYQ KFHIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEDNE NFQRRCYWML 120 EQELMHIVFV HYLEVKGNRI SSSGTKENNS NSLSGSTSVN IESTANTSST LSPLCEDADS 180 DHLALLKVLP LLFLGNRDGW ASQVHVNRVK ESDSQRLVGI PTLDASFENP LARFQTLPPY 240 NNALLAQTNP SNTGLMSVEG HLRSPLQNQV NWQIPVQDSL PLQKWPPMDS HSGMAAHGTD 300 LTLHETFSSL IGSQNQQPSF QAPFTSVEAA YIPKFGPEDL LYEASANQTL PLRKSLLKKE 360 DSLKKVDSFS RWVSNELAEM EDLQMQSSSG GGIAWTSVET AAAASSLSPS LSKDQRFTMI 420 DFWPKWTQTD SEVEVMVIGT FLLSPQEVTS YSWACMFGEV EVPAEILVDG VLCCHAPPHE 480 VGQVPFYITC SDRFSCSEVR EFDFLPGSTR KLNAADLYGA FINEASLHLR FENLLARVSS 540 SAQEHSIFED VGEKRRKISR IMLLKDEKES LLASTTEKGL TEVEAKERLI REEFEDKLYL 600 WLIHKVTEEG KGANILDEEG QGVLHLAAAL GYDWAIKPIL AAGVSINFRD ANGWEDTVAL 660 LVSLGADSGA LTDPSPELPL GKTASDLAYG NGHRGISGFL AESSLTSYLE KLTVDGKEDS 720 STDSSRAKAV QTVAERTATP MSYGDVPETL SMKDSLTAVL NATQAADRLH QVFRMQSFQR 780 KQLSEIGEKN EFGVSDELAV SFAAGKSKKA GHSSGGAAVH AAAVQIQKKY RGWKKRKEFL 840 LIRQRIVKIQ AHVRGHQVRK QYRAIIWSVG LLEKIILRWR RKGSGLRGFK RDAITKAPEP 900 VCAAPAQEDD YDFLKEGRKQ TEERLQKALT RVKSMAQYPE ARAQYRRLLT VVEGIRENEA 960 SSSSAMNNNN NNNISNTEEA ANYNEEDDLI DIDSLLDDDT FMSLAFE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
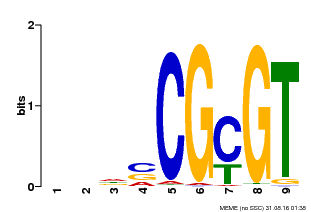
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018456913.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A397XS62 | 0.0 | A0A397XS62_BRACM; Uncharacterized protein | ||||
| STRING | Bo9g018010.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



