 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC1269_p4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 349aa MW: 38185.7 Da PI: 7.8417 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.6 | 0.0004 | 136 | 158 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y+C+ C+++F++ L H +H
RrC1269_p4 136 YQCKTCDRTFPSFQALGGHRASH 158
9***********99999998887 PP
| |||||||
| 2 | zf-C2H2 | 12.5 | 0.00045 | 216 | 238 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg F++ L H+r+H
RrC1269_p4 216 HECGVCGAEFTSGQALGGHMRRH 238
78********************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.5E-8 | 135 | 158 | No hit | No description |
| Pfam | PF13912 | 2.0E-12 | 135 | 160 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.028 | 136 | 158 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.471 | 136 | 163 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 138 | 158 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.5E-8 | 211 | 238 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.1E-4 | 214 | 240 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF13912 | 1.5E-11 | 215 | 240 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.054 | 216 | 238 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.951 | 216 | 240 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 218 | 238 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MEAFEEAIAA SKEQSLIFKG KRTKRQRPQS PIPFSISPPI VPSHARDIEE ESKKDGVITS 60 SSSSASWFSN NNATLKAEEE EEEQDIANCL ILLSQGHSLP IPNHEANNNN TYKFSSRRFL 120 ETSSSNGGGK AGYYVYQCKT CDRTFPSFQA LGGHRASHKK PKATLSLYSN IDVKKNIYES 180 EAVSLVTTSA IYNNNNNNNN NRSVAVYGKA GSNKVHECGV CGAEFTSGQA LGGHMRRHRG 240 AVVVAAAPVN IVTVAMAAAN TELSLSSMSF DQISDGQDHL VMPATKRAKK PVVSLDLDLN 300 LPAPEDENRV NGFTFALKQK QEQEHQPTKQ REEPKCLLLS APTLVDCHY |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00487 | DAP | Transfer from AT5G04390 | Download |
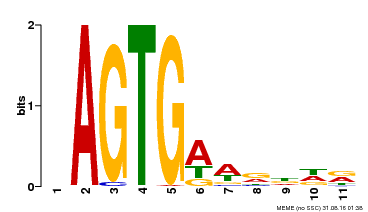
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018444409.1 | 0.0 | PREDICTED: zinc finger protein ZAT5 | ||||
| TrEMBL | A0A3P6EAG5 | 0.0 | A0A3P6EAG5_BRAOL; Uncharacterized protein | ||||
| STRING | Bra009457.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2231 | 27 | 75 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G04390.1 | 1e-156 | C2H2 family protein | ||||



