 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 30174.m008969 | ||||||||
| Common Name | LOC8258519, RCOM_1613930 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 825aa MW: 89458.4 Da PI: 6.1829 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.6e-21 | 123 | 178 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
30174.m008969 123 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 178
688999***********************************************999 PP
| |||||||
| 2 | START | 204.8 | 3.4e-64 | 332 | 556 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
ela++a++elvk+a+ ++p+W +s e++n +e++++f++ + + ea+r+ g+v+ ++ lve+l+d++ +W e+++ + +t++
30174.m008969 332 ELALAAMDELVKMAQTDDPLWIRSLeggrEMLNHEEYVRTFTPCIGmkpsgFVFEASREAGMVIINSLALVETLMDSN-RWAEMFPcviaRTSTTD 426
5899**************************************9998999999**************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE- CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdl 172
vissg g lqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + ++ +++lpSg+++++++ng+skvtwveh+++
30174.m008969 427 VISSGmggtrnGSLQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGGPAFANCRRLPSGCVVQDMPNGYSKVTWVEHAEY 522
***********************************************************999********************************** PP
-SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 173 kgrlphwllrslvksglaegaktwvatlqrqcek 206
+++ +h+l+r+l++sg+ +ga++wvatlqrqce+
30174.m008969 523 DESPIHQLYRPLISSGMGFGAQRWVATLQRQCEC 556
********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.11E-20 | 106 | 180 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.2E-22 | 110 | 180 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 120 | 180 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 121 | 184 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.34E-18 | 122 | 180 | No hit | No description |
| Pfam | PF00046 | 1.4E-18 | 123 | 178 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 155 | 178 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.77 | 323 | 559 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.47E-33 | 325 | 556 | No hit | No description |
| CDD | cd08875 | 2.36E-125 | 327 | 555 | No hit | No description |
| Pfam | PF01852 | 3.5E-56 | 332 | 556 | IPR002913 | START domain |
| SMART | SM00234 | 1.0E-48 | 332 | 556 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.51E-23 | 585 | 818 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 825 aa Download sequence Send to blast |
MSFGGFLENG SPGGGGARIV ADIPFNNNSS SSSTNMPTGA IAQPRLLSPS FTKSMFNSPG 60 LSLALQQPNI DGQGDHVARM AENFETIGGR RSREEEHESR SGSDNMDGAS GDDQDAADNP 120 PRKKRYHRHT PQQIQELEAL FKECPHPDEK QRLELSKRLC LETRQVKFWF QNRRTQMKTQ 180 LERHENSLLR QENDKLRAEN MTIRDAMRNP ICSNCGGPAI IGDISLEEQH LRIENARLKD 240 ELDRVCALAG KFLGRPISSL ASSIGPPMPN SSLELGVGNN GFAGLSTVAT TLPLGPDFGG 300 GISTLNVVTQ TRPGNTGVTG LDRSLERSMF LELALAAMDE LVKMAQTDDP LWIRSLEGGR 360 EMLNHEEYVR TFTPCIGMKP SGFVFEASRE AGMVIINSLA LVETLMDSNR WAEMFPCVIA 420 RTSTTDVISS GMGGTRNGSL QLMHAELQVL SPLVPVREVN FLRFCKQHAE GVWAVVDVSI 480 DTIRETSGGP AFANCRRLPS GCVVQDMPNG YSKVTWVEHA EYDESPIHQL YRPLISSGMG 540 FGAQRWVATL QRQCECLAIL MSSTVPARDH TAAITASGRR SMLKLAQRMT DNFCAGVCAS 600 TVHKWNKLNA GNVDEDVRVM TRKSVDDPGE PPGIVLSAAT SVWLPVSPQR LFDFLRDERL 660 RSEWDILSNG GPMQEMAHIA KGQDHGNCVS LLRASAMNAN QSSMLILQET CIDAAGSLVV 720 YAPVDIPAMH VVMNGGDSAY VALLPSGFAI VPDGPGSRGS PTNQNGGGNN GGGPNRVSGS 780 LLTVAFQILV NSLPTAKLTV ESVETVNNLI SCTVQKIKAA LQCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
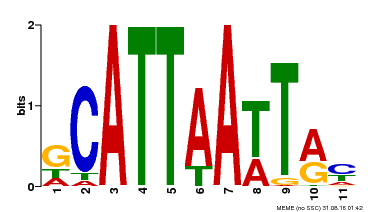
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 30174.m008969 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002511801.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | B9RDL2 | 0.0 | B9RDL2_RICCO; Homeobox protein, putative | ||||
| STRING | XP_002511801.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 30174.m008969 |
| Entrez Gene | 8258519 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



