 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 29686.m000890 | ||||||||
| Common Name | LOC8276332, RCOM_0622120 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 556aa MW: 60406.2 Da PI: 4.8728 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.5 | 6.4e-18 | 40 | 87 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+++vk++G g+W+++ ++ g+ R++k+c++rw ++l
29686.m000890 40 KGPWTSAEDAILIEYVKKHGEGNWNAVQKHSGLSRCGKSCRLRWANHL 87
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 53.9 | 4.1e-17 | 93 | 136 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE++l++++++++G++ W++ a++++ gRt++++k++w++
29686.m000890 93 KGAFTQEEEQLIIELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 136
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.689 | 35 | 87 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.24E-31 | 39 | 134 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.6E-16 | 39 | 89 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-16 | 40 | 87 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.7E-25 | 41 | 94 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.41E-12 | 42 | 87 | No hit | No description |
| PROSITE profile | PS51294 | 26.872 | 88 | 142 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.3E-17 | 92 | 140 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.4E-16 | 93 | 136 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-26 | 95 | 141 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.11E-12 | 95 | 136 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 556 aa Download sequence Send to blast |
MSHTTNESDD GLFSKDRIDS PLAEGGNCGG SANGGAMLKK GPWTSAEDAI LIEYVKKHGE 60 GNWNAVQKHS GLSRCGKSCR LRWANHLRPN LKKGAFTQEE EQLIIELHAK MGNKWARMAA 120 HLPGRTDNEI KNYWNTRIKR RQRAGLPLYP PEVSFQALQE SHQGLTIGGI NTGDKVHGDL 180 LRNNGYEIPD VIFDSLKASQ GISPYVPELA DITTSSMLIK GLSSSQYGSF MSPSAHRQKR 240 LRESTTFIPG YSGNIKSEFP LFDQFQDESC DKVAQSFGLS FPFDPDPTKN PQSFGENQGS 300 HTLANGDFST SKLTSGPVKL ELPSLQYPET DLGSWGTSPP PLLETLDNFI QSPPTAIIES 360 SPRNSGLLDA LLYEAKALSS AKNHSSEKSS NSSTVTPGEL AECSAINICE TEWEDYGDPL 420 SPLGHTATSL FSECTPISAS GSSLDEPPPT DTFTGCIVKT ELVSQPWSPE RDKETTTRVD 480 ITRPDALLAS DWLGQGSGYI KDQVIMTDNI ATLLGDDLSS DYKQMSAGAS TSNQVWGLGS 540 CAWNNMPAVC QMSELP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-32 | 38 | 141 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
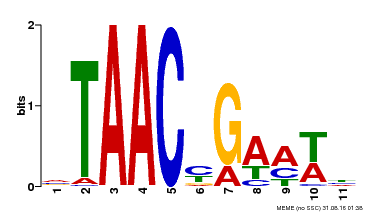
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 29686.m000890 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002525574.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_015578733.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_015578734.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_015578735.1 | 0.0 | transcription factor MYB33 | ||||
| TrEMBL | B9SHV5 | 0.0 | B9SHV5_RICCO; R2r3-myb transcription factor, putative | ||||
| STRING | XP_002525574.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-141 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 29686.m000890 |
| Entrez Gene | 8276332 |



