 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 29216.m000261 | ||||||||
| Common Name | RCOM_0405730 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 355aa MW: 38981.6 Da PI: 9.7882 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.7 | 8.6e-13 | 283 | 328 | 6 | 55 |
HHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 6 nerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+++Er RR ri +++ +L+el+P+ +k + a++L +Av+YIk+Lq
29216.m000261 283 SIAERVRRTRISERMRKLQELVPNM----DKQTNTADMLDLAVDYIKDLQ 328
689*********************7....688*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 15.25 | 277 | 327 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 5.76E-16 | 280 | 339 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.75E-12 | 281 | 330 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.0E-14 | 282 | 339 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.4E-13 | 283 | 333 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 9.4E-10 | 283 | 328 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 355 aa Download sequence Send to blast |
MDSSSNPNFH HQNNQQPSSG LLRFRSAPSS FLANFNDNGV TSNNSVMGFQ ELEDKSAVRV 60 REVGAVVNYA NSTQSYSGLP PHYPRHTNSS AATSSSAMDS SFGLIGSMAM GGHHEQEAKR 120 VDSNLARQSS SPAGLFGNLS VQTGYATMKG MGNYARVNGT SGEVSPRLKS QLSFPSRIPS 180 SLSMLSQISE IGSESIGFPY GSWNDSPFSE NFNGMKRDPD DNGKPFSAAQ NGELGNRVHL 240 LSHHLSLPKA SVDMVAMEKF LHFQDSVPCK IRAKRGCATH PRSIAERVRR TRISERMRKL 300 QELVPNMDKQ TNTADMLDLA VDYIKDLQKQ YKTLSDNRAN CKCLSKQKPV QNQIV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00308 | DAP | Transfer from AT2G42280 | Download |
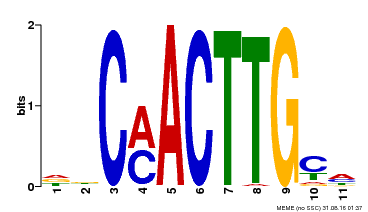
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 29216.m000261 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015582852.1 | 0.0 | transcription factor bHLH130 isoform X1 | ||||
| Swissprot | Q66GR3 | 1e-114 | BH130_ARATH; Transcription factor bHLH130 | ||||
| TrEMBL | B9T2D6 | 0.0 | B9T2D6_RICCO; DNA binding protein, putative | ||||
| STRING | XP_002532405.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5177 | 32 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42280.1 | 1e-110 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 29216.m000261 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



