 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 28883.m000751 | ||||||||
| Common Name | LOC8260405, RCOM_0353920 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 393aa MW: 43512.7 Da PI: 6.4 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.5 | 2.6e-32 | 233 | 288 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs+ AtPk+i+elmkv+gLt+++vkSHLQkYRl+
28883.m000751 233 KQRRCWSPELHRRFLHALQQLGGSHAATPKQIRELMKVDGLTNDEVKSHLQKYRLH 288
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.466 | 230 | 290 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-29 | 230 | 291 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.06E-18 | 231 | 291 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.0E-27 | 233 | 288 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.6E-7 | 235 | 286 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 393 aa Download sequence Send to blast |
MDYAEKMQRC HEYVEALEEE KRKIQVFQRE LPLCLELVTQ AIEACKRELS GTTTEYMHGQ 60 SECSEQTTST DGTANGTGTR SLVLEEFIPI KRINSSSHND NDNDDDNENE KEDNDDDEEE 120 EDQDSHKPNK SIRDINNDQK KKSDWLRSVQ LWNQSSPDSE PPKEDLPRKA AVTEVKRNGG 180 AFQPFHKEKG IAKTPPSVPA SATSSSAETG TGGGTSGAGN NRKEDKDGQA QRKQRRCWSP 240 ELHRRFLHAL QQLGGSHAAT PKQIRELMKV DGLTNDEVKS HLQKYRLHTR RPSPTIHNNS 300 NPQAPQFVVV GGIWVPPPEY AAVAATTASM ETVTTAAANG IYAPVAAPLG TIPKQQRAQS 360 QHLQSERRGS HSERSNSPAT SSSTHTTTNS PVF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 5e-13 | 233 | 286 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-13 | 233 | 286 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-13 | 233 | 286 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-13 | 233 | 286 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-13 | 233 | 286 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
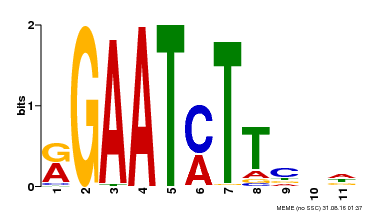
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 28883.m000751 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002529155.1 | 0.0 | transcription factor HHO3 isoform X2 | ||||
| Swissprot | Q9FPE8 | 1e-113 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | B9ST36 | 0.0 | B9ST36_RICCO; DNA binding protein, putative | ||||
| STRING | XP_002529155.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3918 | 33 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-107 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 28883.m000751 |
| Entrez Gene | 8260405 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



