 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.011G191300.1 | ||||||||
| Common Name | PHAVU_011G191300g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 555aa MW: 61028.7 Da PI: 5.2281 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.6 | 5.9e-18 | 37 | 84 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed++lvd+v+++G g+W+++ ++ g+ R++k+c++rw ++l
Phvul.011G191300.1 37 KGPWTSTEDDILVDYVNKHGEGNWNAVQKHTGLLRCGKSCRLRWANHL 84
79******************************99**********9986 PP
| |||||||
| 2 | Myb_DNA-binding | 48.4 | 2.2e-15 | 90 | 133 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ ++++++++G++ W++ a++++ gRt++++k++w++
Phvul.011G191300.1 90 KGAFTAEEERFIAELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 133
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.574 | 32 | 84 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.4E-14 | 36 | 86 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.42E-29 | 36 | 131 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 4.1E-16 | 37 | 84 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-23 | 38 | 90 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.45E-11 | 39 | 84 | No hit | No description |
| PROSITE profile | PS51294 | 24.53 | 85 | 139 | IPR017930 | Myb domain |
| SMART | SM00717 | 8.5E-16 | 89 | 137 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-14 | 90 | 133 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-24 | 91 | 138 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.42E-11 | 92 | 133 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 555 aa Download sequence Send to blast |
MRRVSNEIED EVLPSDLSAA QLNDESYEGS TSIVLKKGPW TSTEDDILVD YVNKHGEGNW 60 NAVQKHTGLL RCGKSCRLRW ANHLRPNLKK GAFTAEEERF IAELHAKMGN KWARMAAHLP 120 GRTDNEIKNY WNTRIKRRQR AGLPLYPPEV CLQGFQESQR NQSNGGLNGG DKMHLDLLQT 180 NSYEIHGAIF DSLKDNQGIL PYAPKFPDAS VYGNMLKGLD SSQYCSVMTP ASPKHKRLRE 240 STIPFIGSND TRKNGLYPFS QIRDNNSDKI AQSFGMQALL DPGPSSHNSM CYSHSLSNGN 300 SSTSKPTFEA VKLELPSLQY PELDLGSWGT SPPPPLLESV DDFFQSPTPI STLESDCSSP 360 QNSGLLDALL YQAKTLSSSK NHCSDKSSNS STATPGDRGD SSALNVYETE WEDYADPVSP 420 FGATSILNEC PALLRASENS LDEQLSFQAF TGNIPKLEFV DQVWTPESEN QTLSLLNFTR 480 PDFLFASDWH DLGSGNGKNQ AITTDTTMTL LGDHFASDRK HMTSGTSQSS QVWGFGSCAT 540 ANNMPALCQL SDLG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-31 | 35 | 138 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
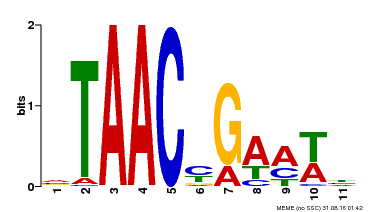
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.011G191300.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC525897 | 0.0 | KC525897.1 Glycine max GAMBY1-like family protein (GAMYB1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007133583.1 | 0.0 | hypothetical protein PHAVU_011G191300g | ||||
| TrEMBL | V7AIZ8 | 0.0 | V7AIZ8_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007133583.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-117 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.011G191300.1 |
| Entrez Gene | 18616680 |



