 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.002G328700.2 | ||||||||
| Common Name | PHAVU_002G328700g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 308aa MW: 34817.1 Da PI: 6.5842 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 105.7 | 5.6e-33 | 63 | 203 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkk......lelee...vikev.diykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratk.. 79
lp+G++F+P+d+e++ e+L++kv + ++ e +++ +i++++P++Lp v+++ + +fF++ +k+y+tg+rk+r+++
Phvul.002G328700.2 63 LPAGVKFDPNDQEIL-EHLEAKVLCDVpklhplID--EfipTLEGEnGICYTHPEKLP-GVSKDGQIRHFFHRPSKAYTTGTRKRRKVHtd 149
799***********9.99****9998856643322..22222333336**********.555666777******************74444 PP
NAM 80 ....sgyWkatgkdkevlskkgelvglkktLvfy..kgrapkgektdWvmheyrl 128
+ +W++tgk+++v++ +g+++g kk+Lv+y gr++k+ekt+Wvmh+y+l
Phvul.002G328700.2 150 eegsETRWHKTGKTRPVFV-GGAVKGFKKILVLYtnYGRQKKPEKTNWVMHQYHL 203
6555778************.9*************6658***************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 8.63E-38 | 58 | 222 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 33.352 | 63 | 222 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.2E-14 | 64 | 203 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 308 aa Download sequence Send to blast |
MTWCNDSDEE RGIELLAPTL TPTTNNTLLT DTKPSENRTI TCPSCGHNIE FQDQTGINDL 60 PGLPAGVKFD PNDQEILEHL EAKVLCDVPK LHPLIDEFIP TLEGENGICY THPEKLPGVS 120 KDGQIRHFFH RPSKAYTTGT RKRRKVHTDE EGSETRWHKT GKTRPVFVGG AVKGFKKILV 180 LYTNYGRQKK PEKTNWVMHQ YHLGSNEEEK DGELVVSKVF YQTQPRQCGN SITKQDLYQS 240 VDDEDIIPRS GTLMEYYNPS FINYDHVGGR NNREASPQLI PNLVMQADTS SFIRLAMDAT 300 KARLDTK* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that plays a regulatory role in the development of secondary cell wall fibers (PubMed:18952777,PubMed:22133261). Involved in the regulation of cellulose and hemicellulose biosynthetic genes as well as of genes involved in lignin polymerization and signaling (PubMed:22133261). Is not a direct target of SND1 (PubMed:18952777). {ECO:0000269|PubMed:18952777, ECO:0000269|PubMed:22133261}. | |||||
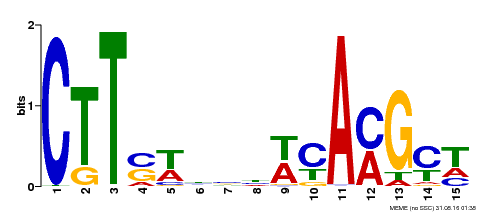
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00459 | DAP | Transfer from AT4G28500 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.002G328700.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 1e-147 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007160521.1 | 0.0 | hypothetical protein PHAVU_002G328700g | ||||
| Refseq | XP_007160522.1 | 0.0 | hypothetical protein PHAVU_002G328700g | ||||
| Swissprot | O49459 | 1e-123 | NAC73_ARATH; NAC domain-containing protein 73 | ||||
| TrEMBL | V7CUD5 | 0.0 | V7CUD5_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007160521.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G28500.1 | 1e-122 | NAC domain containing protein 73 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.002G328700.2 |
| Entrez Gene | 18639855 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



