 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.002G282200.1 | ||||||||
| Common Name | PHAVU_002G282200g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 843aa MW: 93474.7 Da PI: 6.6 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.8 | 5e-24 | 159 | 260 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+lt sd++++g +++ +++a+e+ + k++ +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F
Phvul.002G282200.1 159 FCKTLTASDTSTHGGFSVLRRHADEClppldmS-KQPPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL-- 246
99*****************************63.4445569************************************************.. PP
SSSEE..EEEEE-S CS
B3 86 grsefelvvkvfrk 99
+ +++el+v+v+r+
Phvul.002G282200.1 247 RGENGELRVGVRRA 260
449999*****996 PP
| |||||||
| 2 | Auxin_resp | 119.7 | 2.3e-39 | 285 | 367 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+ t+++F+v+Y+Pr+s++eF+v++++++++lk+++++GmRfkm+fe+e+++e+r++Gt+vg++d+dp rWpnSkWrsLk
Phvul.002G282200.1 285 AWHAILTGTMFTVYYKPRTSPAEFIVPYDQYMESLKNNYTIGMRFKMRFEGEEAPEQRFTGTIVGIEDADPKRWPNSKWRSLK 367
79*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.28E-40 | 155 | 288 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 9.9E-42 | 155 | 274 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.59E-22 | 157 | 259 | No hit | No description |
| PROSITE profile | PS50863 | 12.026 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.6E-22 | 159 | 260 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.8E-23 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.8E-37 | 285 | 367 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 24.664 | 719 | 801 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 4.91E-11 | 720 | 796 | No hit | No description |
| Pfam | PF02309 | 2.7E-8 | 765 | 807 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MASSEVSIKG NLVNGKGENS SGGFNNGNDV RNAGGEPQNG SSSSSARDAE TALYRELWHA 60 CAGPLVTVPR EGERVFYFPQ GHIEQVEAST NQVAEQHMPV YDLPPKILCR VINVMLKAEP 120 DTDEVFAQVT LLPEPNQDEN AVEKEGPPAP PPRFHVHSFC KTLTASDTST HGGFSVLRRH 180 ADECLPPLDM SKQPPTQELV AKDLHGNEWR FRHIFRGQPR RHLLQSGWSV FVSSKRLVAG 240 DAFIFLRGEN GELRVGVRRA MRQQGNVPSS VISSHSMHLG VLATAWHAIL TGTMFTVYYK 300 PRTSPAEFIV PYDQYMESLK NNYTIGMRFK MRFEGEEAPE QRFTGTIVGI EDADPKRWPN 360 SKWRSLKVRW DETSNVPRPE RVSQWKIEPA LAPPALNPLP MPRPKRPRSN VVPSSPDSSV 420 LTREASSKVS VDPLPASGFQ RVLQGQELST LRVNFAESNE SDTAEKSAWS SAADDEKIDV 480 VSTSRRYGSE SWMSMGRHEP TYPDLLSGFG VHGDQSSHPS FVDQNGPVAN LSRKHFLDRE 540 GKHNVLSPWP SLPLNLLDSN TKASAQGGDT TCQVRGNMRF SSAFGDYTVL HGHKVEHSHG 600 NFLMPPPLST QYESPRSREL LPKPISGKPK DSDCKLFGIS LLSSPIVLDP SVSQRNVAIE 660 PVGHMHNQQH TFENDTKSEN SRGLKPADGL LIDDHEKLSQ NSQPHLKDVQ PKSNSGSARS 720 CTKVHKKGIA LGRSVDLTKF SAYDELIAEL DQLFEFGGEL TSPQKDWLIV YTDNEGDMML 780 VGDDPWQEFV AMVRKIYIYP KEEIQKMSPG TLSSKNEENQ SVSEGVEAQE IKCQLNHSAS 840 EA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-178 | 52 | 389 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-178 | 52 | 389 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-178 | 52 | 389 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
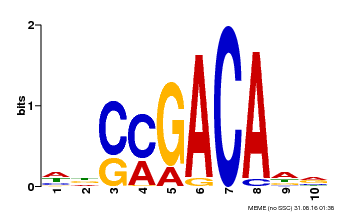
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.002G282200.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007159966.1 | 0.0 | hypothetical protein PHAVU_002G282200g | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | V7CRR6 | 0.0 | V7CRR6_PHAVU; Auxin response factor | ||||
| STRING | XP_007159966.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.002G282200.1 |
| Entrez Gene | 18639375 |



