 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.001G258000.2 | ||||||||
| Common Name | PHAVU_001G258000g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 485aa MW: 54533.3 Da PI: 5.9217 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 77.4 | 1.9e-24 | 128 | 181 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++W+ +LH++Fv+av+q+ G +k Pk+il+lm+v+ Lt+e+v+SHLQkYRl
Phvul.001G258000.2 128 KARVVWSVDLHQKFVKAVNQI-GFDKVGPKKILDLMNVPWLTRENVASHLQKYRL 181
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 32.1 | 5.8e-12 | 1 | 56 | 53 | 109 |
CTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 53 mpgmdGlellkeireeepklpiivvtahgeeedalealkaGakdflsKpfdpeelvk 109
mp+mdG++ll++ e +lp+i+++ ge + + + ++ Ga d+l Kp+ ++el++
Phvul.001G258000.2 1 MPDMDGFKLLEHVGLEM-DLPVIMMSVDGETSRVMKGVQHGACDYLLKPIRMKELRN 56
9************6644.8***********************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 9.5E-21 | 1 | 83 | No hit | No description |
| CDD | cd00156 | 3.33E-12 | 1 | 61 | No hit | No description |
| PROSITE profile | PS50110 | 23.493 | 1 | 62 | IPR001789 | Signal transduction response regulator, receiver domain |
| SuperFamily | SSF52172 | 2.99E-15 | 1 | 74 | IPR011006 | CheY-like superfamily |
| Pfam | PF00072 | 6.6E-9 | 1 | 57 | IPR001789 | Signal transduction response regulator, receiver domain |
| SuperFamily | SSF46689 | 4.75E-17 | 126 | 183 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-27 | 126 | 185 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-21 | 128 | 181 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 130 | 180 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 485 aa Download sequence Send to blast |
MPDMDGFKLL EHVGLEMDLP VIMMSVDGET SRVMKGVQHG ACDYLLKPIR MKELRNIWQH 60 VYRKRIHEAK EFESFESFES IHLMTNGSEL SDEGNLFAVE DVTSTKKRKD ADNKHDDKEC 120 MDPSSTKKAR VVWSVDLHQK FVKAVNQIGF DKVGPKKILD LMNVPWLTRE NVASHLQKYR 180 LYLIRIQKEN DRRSSSSGMK HSDFPSKDLG SFGFQNPVNK QQNDVSIDSY NYSDGSLQLQ 240 NVENKSHEGD KGTVSQFTIA KKGRPLRESS RVGLQQPFDC SMPTQYSWTE VPHTQLKEEH 300 KSLVHLEDSF NQMPLQGKQH PIQVDQSQSV ASISSVPSIR EEGVAACIEA KPLFSDFKND 360 QSSSVNSIGN AVDTFPIQPG SLVMNAQSLA SSTTNSGLKA HGYNNLSCIS DLEIYQRNLL 420 LGASAPLDED LHFHWLQGEC YNMNFGLQNI GMPEYYDPGL IAEAPSHLYD SSDYSVMDQG 480 LFIA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-17 | 124 | 186 | 1 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
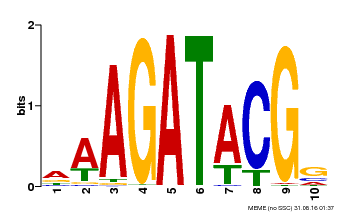
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.001G258000.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 0.0 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007163715.1 | 0.0 | hypothetical protein PHAVU_001G258000g | ||||
| TrEMBL | V7D026 | 0.0 | V7D026_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007163714.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 9e-83 | response regulator 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.001G258000.2 |
| Entrez Gene | 18642570 |



